
Future Directions
Workshop: Advancing
the Next Scientific
Revolution in
Toxicology
April 28-29,2022
Thomas Hartung, Johns Hopkins University, University of Konstanz,
and Georgetown University
Ana Navas-Acien, Columbia University
Weihsueh A. Chiu, Texas A&M University
Prepared by:
Kate Klemic, Virginia Tech Applied Research Corporation
Matthew Peters, Virginia Tech Applied Research Corporation
Shanni Silberberg, Ofce of the Under Secretary of Defense
(Research & Engineering), Basic Research Ofce
Future Directions Workshop series
Workshop sponsored by the Basic Research Ofce, Ofce of
the Under Secretary of Defense for Research & Engineering
CLEARED
For Open Publication
Department of Defense
OFFICE OF PREPUBLICATION AND SECURITY REVIEW
Apr 26, 2023
ii
Contents
Preface iii
Executive Summary 1
Introduction 4
Toxicology Research Challenges 6
Toxicology Research Advances and Opportunities 8
Toxicology Research Trajectory 15
Accelerating Progress 19
Conclusion 20
Glossary 21
Bibliography 22
Appendix I—Workshop Attendees 27
Appendix II—Workshop Agenda and Prospectus 34

iii
Innovation is the key
to the future, but basic
research is the key to
future innovation.
– Jerome Isaac Friedman,
Nobel Prize Recipient (1990)
Preface
Over the past century, science and technology has brought
remarkable new capabilities to all sectors of the economy,
from telecommunications, energy, and electronics to medicine,
transportation and defense. Technologies that were fantasy
decades ago, such as the internet and mobile devices, now
inform the way we live, work, and interact with our environment.
Key to this technological progress is the capacity of the global
basic research community to create new knowledge and to
develop new insights in science, technology, and engineering.
Understanding the trajectories of this fundamental research,
within the context of global challenges, empowers stakeholders
to identify and seize potential opportunities.
The Future Directions Workshop series, sponsored by the
Basic Research Directorate of the Ofce of the Under Secretary
of Defense for Research and Engineering, seeks to examine
emerging research and engineering areas that are most likely to
transform future technology capabilities. These workshops gather
distinguished academic researchers from around the globe
to engage in an interactive dialogue about the promises and
challenges of each emerging basic research area and how they
could impact future capabilities. Chaired by leaders in the eld,
these workshops encourage unfettered considerations of the
prospects of fundamental science areas from the most talented
minds in the research community.
Reports from the Future Direction Workshop series capture
these discussions and therefore play a vital role in the discussion
of basic research priorities. In each report, participants are
challenged to address the following important questions:
• How will the research impact science and technology
capabilities of the future?
• What is the trajectory of scientic achievement over the next
few decades?
• What are the most fundamental challenges to progress?
This report is the product of a workshop held April 28-29, 2022, at
the Basic Research Innovation Collaboration Center in Arlington,
VA on the future of toxicology research. It is intended as a
resource for the S&T community including the broader federal
funding community, federal laboratories, domestic industrial
base, and academia.

1
Executive Summary
1 hps://pubmed.ncbi.nlm.nih.gov/20574894/
In the nearly two decades since the human genome
was sequenced, the eld of toxicology has undergone a
transformation, taking advantage of the explosion in biomedical
knowledge and technologies to move from a largely empirical
science aimed at ensuring the absence of harmful effects to a
mechanistic endeavor aimed at elucidating disease etiology
based on an understanding of the biological responses to
chemicals (including biochemistry) and the impact on organ
systems. However, a substantial gap remains between the
promise of mechanistic toxicology and its actual impacts on
improving human health. Toxicology continues to work in a
largely reductionist paradigm of single endpoints, chemicals,
and biological targets, whereas it is known that biology and
pathobiology involve complex interactions across each of these,
with the additional recognition that social stressors also have
biological consequences. At the same time, the pace of scientic
and technical advances has resulted in a deluge of models and
data for understanding toxicological exposure, hazard, and
risk that is increasingly challenging to evaluate, integrate and
interpret. A critical need, therefore, exists to understand how to
leverage these new frontiers in toxicology to achieve the desired
long-term impact of improving human health. This fundamental
problem addresses the question of what exposures, now or
in the future, can contribute to disease and calls for a Human
Exposome Project.
The 2007 National Research Council report on Toxicity Testing
for the 21st Century—a Vision and a Strategy
1
(Tox-21c) was a
watershed moment for US toxicology, changing the discussion
from whether to change to when and how to change. With
knowledge in the life sciences doubling every seven years since
1980 and every 3.5 years since 2010, as well as publications
doubling every fteen years (Bornmann, 2021; Densen, 2011), we
now have about 16 times as much knowledge and twice as many
publications as in 2007.
The Future Directions in Toxicology Workshop convened on April
28-29, 2022, in Arlington, VA, to examine research challenges
and opportunities to usher toxicology into a new paradigm
as a predictive science. Hosted by the Basic Research Ofce
in the Ofce of the Under Secretary of Defense for Research
and Engineering, this workshop gathered 20 distinguished
researchers from across academia, industry, and government
to discuss how basic research can advance the science of
toxicology. The workshop aimed at the next generation of a
vision for toxicology, “Toxicity Testing for the 21st Century
2.0—Implementation” that extends the vision of the 2007 report
and adapts it to scientic and technological progress. This report
is the product of those discussions, summarizing current research
challenges, opportunities, and the trajectory of toxicological
science for the next twenty years.
The vision developed at the workshop foresees toxicology
developing into a Human Exposome Project that better
integrates the exposure side of disease, focusing on real-
world exposures affecting diverse populations over time. Thus,
changing the principal approach from a hazard-driven to an
exposure-driven paradigm. This new paradigm identies the
relevant human or ecological exposures and then bases the
risk assessment process on exposomics, forming an exposure/
mechanism hypothesis from the multi-omics imprint in biouids
and tissues, and biomonitoring, the large-scale sampling
and measurement of biospecimen. This new paradigm also
incorporates negligible exposures and is focused on ensuring
safety instead of predicting toxicity. Another critical aspect of this
paradigm is the inclusion of disruptive research technologies,
such as microphysiological systems, the bioengineering of
organ architecture and functionality to model (patho-)physiology,
and articial intelligence/machine learning to process the
complex data generated for informed decisions. Ultimately, we
need to integrate the evidence provided by these technologies,
especially through probabilistic risk assessment. An evidence-
based toxicology approach ensures condence and trust in the
process by which scientic evidence is assessed for the safety of
chemicals for human health and the environment.
The workshop was organized around three key areas that are
likely to transform toxicology: 1) employing an exposure-driven
approach, 2) utilizing technology-enabled techniques, and 3)
embracing broad-scale evidence integration. These three key
areas are expected to have a huge impact on the development
of three key long-term public health goals: 1) Precision Health,
2) Targeted Public Health Interventions and Environmental
Regulations and 3) Safer Drugs and Chemicals, through their
distinct perspectives and long-term goals.
Exposure-driven Toxicology
Exposure-driven assessments were not covered in the 2007
Tox-21c report and were only the subject of a parallel NRC
report, but the needs for integration into toxicology, for example
through exposomics, are increasingly evident. Exposure-
driven toxicology, focused on real-world exposures and gene-
environment interactions that affect diverse populations can
contribute to addressing the three aims identied during the
workshop: 1) precision health through the identication of
environmental exposures for improved health outcomes in
specic populations, 2) targeted public health interventions and
environmental regulations to address those environmentally-
driven health outcomes and 3) the identication of safer drugs
and chemicals. Precision health aims for individual, personalized
preventive interventions, and pharmaceutical and non
pharmaceutical therapies. Targeted public health Interventions
and environmental regulations must address population and
spatial-temporal variability in genome and epigenome, as well as
exposome. Safer drugs and chemicals shall be attained through
in vitro/in silico chemical screening, in vitro/in silico clinical trials
and identifying intrinsic and extrinsic susceptibilities.

2
Workshop participants envision a Tox-21c 2.0 that reects real-
world based exposure designs (in silico, cellular, organoids,
models, organisms, longitudinal epidemiological studies). It will
include population-scale measurements that are based on readily
available biobanks and ecobanks that inform on the distribution
of thousands of chemical and non-chemical stressors in relevant
populations (general population, relevant subgroups, disease
cohorts). Noteworthy, the exposomics approach potentially
can interrogate all types of stressors, not just chemicals, that
actually perturb biology and change biomarkers in body uids.
Study designs and computational approaches will be aligned to
provide interpretable and actionable results. Ethical issues, policy
implications, community engagement, and citizen participation
will keep pace with and inform the technology, rather than being
exclusively reactive to the technology. In the near-term, a critical
rst implementation step for exposure-driven toxicology and
precision health is to scale-up mass spectrometry technology for
high-quality inexpensive assessment of thousands of chemicals
that can be tagged to exogenous exposures including non-
chemical stressors. Libraries that tag key information for those
chemicals (metadata layering) will need to be expanded and
developed to facilitate interpretation, and to guide preventive
strategies, interventions, and policy recommendations. In the
mid-term, technologies will be required that link the exposome
with health outcomes, and leverage longitudinal studies and
biobanks retrospectively and prospectively, ensuring “FAIR”-ness
(Findability, Accessibility, Interoperability, and Reuse of digital
assets)
2
. In the long-term (20 years), we envision that exposome-
disease predictions and exposome-targeted prevention, and
treatment solutions will become part of the toxicology and
public health practice landscape, leveraging also other ~omics
technologies, genomic information, and clinical characteristics.
Technology-enabled Toxicology
The workshop participants discussed technological advances
over the last 10-15 years of great relevance to toxicology in
three key areas: cell and tissue biology, bioengineering, and
computational methods. Workshop participants noted that
while biological technologies, such as stem cell engineering,
have emerged as routine, commercial enterprises for
biomedical research, their potential in toxicology could be
further expanded through a) reliable and genetically-diverse
cell sourcing, b) improved protocols to differentiate patient-
derived stem cells into adult cell phenotypes across essential
tissues, c) integrative and non-invasive biomarkers, d) integration
of dynamic physiology and pathophysiology outcomes, e)
population heterogeneity and susceptibility through life-courses,
and f) biological surrogates for non-chemical stressors. On
the bioengineering side, workshop participants noted that
technological capabilities, such as microphysiological systems
(MPS), have shown many successes in the laboratory but need
to be further developed to 1) include a variety of models of
increasing architectural complexity (monolayer/suspension
cultures, organoids and multi-organoid systems) for different
stages of drug/chemical development, 2) better represent
healthy and diseased populations by a personalized multiverse
2 hps://www.go-fair.org/fair-principles/
of possible futures, 3) codify platform standardization, 4)
increase throughput, 5) demonstrate validation against in vivo
outcomes, 6) incorporate perfusion and biosensors with near
real-time outputs, and 7) develop automated fabrication. In
addition, the workshop participants noted that the emergence
of “big data” and “big compute” has revolutionized much of
biology, through the ability to analyze and interpret complex and
multi-dimensional information. Computational capabilities and
models are of utmost importance for toxicology, serving as the
key enabling technology. For instance, AI/Machine Learning has
emerged as a key technology to support data mining, predictive
modeling, hypothesis generation, and evidence interpretation
(e.g., explainable AI). Data acquisition and data-sharing following
the FAIR principles is key to unleashing these opportunities. The
emergence of these needs in toxicology necessitates widespread
use and understanding of these technologies combining
them with expert knowledge to yield augmented intelligence
workows. Moreover, given the quantity of information generated
and consumed by these new technologies, the workshop
participants agreed that there is a need for comparable,
compatible, integrable multi-omic databases, quantitative in vitro
to in vivo extrapolation, and the development of in silico “digital
twins” of in vitro and in vivo systems.
Evidence-integrated Toxicology
Workshop participants discussed the key challenge of integrating
data and methods (evidence streams) in test strategies,
systematic reviews, and risk assessments. They agree that
evidence-based toxicology and probabilistic risk assessments are
emerging solutions to this challenge. Evidence integration across
evidence streams (epidemiological, animal toxicology, in vitro, in
silico, non-chemical stressors, etc.) is expected to play a key role
in translating evidence into knowledge that can inform decision-
making. The group developed a vision to conduct complex
rapid/real-time evidence integration by combining advancements
made in data-sharing, and application of articial intelligence
(e.g., natural language processing), with the transparency
and rigor of systematic reviews. To implement this vision, the
workshop participants identied a need for collaborative, open
platform(s) to transparently collect, process, share, and interpret
data, information, and knowledge on chemical and non-chemical
stressors. Creating these platforms is foundational for rapid and
real-time evidence integration and will empower all steps of
protection of human health and the environment. Several needs
were identied to create this platform: 1) software development
to create dynamic and accessible interfaces, 2) denitive
standards and key data elements to facilitate analysis of meta-
data and automated annotation, and 3) consideration for quality
control.
In conclusion, the workshop advocates a paradigm shift
to “Toxicology 2.0” based on the evidence integration of
emerging disruptive technologies, especially exposomics,
microphysiological systems, and machine learning. To date,
exposure considerations typically follow the identication of a
hazard. Future Tox-21c 2.0 must be guided by the identication
3
of relevant exposures through exposomics. The adaptation to
technical progress, especially microphysiological systems and AI,
requires harmonization of reporting and quality assurance. The
key challenge lies in the integration of these different evidence
streams. evidence-based medicine can serve as a role model with
systematic reviews, dened data search strategies, inclusion and
exclusion criteria, risk-of-bias analysis, meta-analysis, and other
evidence synthesis approaches. While this is mostly applicable
to existing data and studies, a new challenge is the prospective
application for the composition of test strategies (Integrated
Test Strategies—ITS, Integrated Approached to Testing and
Assessment—IATA, and Dened Approaches—DA). A key role for
Probabilistic Risk Assessment was also identied. The participants
also emphasized the need to ensure validation of these new
approaches, as well as expand training, communication, and
outreach. Ultimately, it calls for expanding the approach to a
Human Exposome Project.
4
Introduction
In 1983, the US National Research Council (NRC) Committee
on the Institutional Means for Assessment of Risks to Public
Health published a foundational study titled “Risk Assessment
in the Federal Government: Managing the Process” (NRC,
1983), commonly referred to as the Redbook. As Lynn
Goldman put it, the Redbook “has created a framework for
incorporation of toxicology into environmental decision-making
that has withstood the test of time” (Goldman, 2003). It was
complemented by the 2009 report “Science and Decisions:
Advancing Risk Assessment,” aka the Silverbook (NASEM,
2009), which highlighted some challenges in the process.
Parallel work by another NRC committee resulted in the 2007
NRC report on “Toxicity Testing in the 21st Century” (NRC,
2007), or Tox 21c, which developed “a vision and a strategy” to
transform toxicological sciences. The gap analysis of the report
has not really changed, as toxicology is still “time-consuming
and resource-intensive, it has had difculty in meeting many
challenges encountered today, such as evaluating various life
stages, numerous health outcomes, and large numbers of
untested chemicals” and needs “to use fewer animals and
cause minimal suffering in the animals used”. The 2007 report
was complemented by the NRC reports “Exposure Science in
the Twenty-rst Century: A Vision and a Strategy” (NRC, 2012)
and NASEM (2017a) “Using 21st Century Science to Improve
Risk-Related Evaluations.” The Tox-21c and subsequent
reports have changed the debate about safety and risk
assessment of substances in the US and beyond, and led to a
remarkable number of initiatives and programs (Krewski 2020).
The resulting diversity in approaches combined with an ever-
accelerating availability of disruptive technologies calls for a
re-conceptualization of the future of toxicology. The way forward
is to dissolve the dichotomy of hazard and exposure sciences,
embrace the disruptive technological advances, and foster
evidence integration from these evidence streams.
In the nearly two decades since the human genome
was sequenced, the eld of toxicology has undergone a
transformation, taking advantage of the explosion in biomedical
knowledge and technologies to move from a largely empirical
science aimed at ensuring the absence of harmful effects to a
mechanistic endeavor aimed at elucidating disease etiology
and biological response pathways induced by exposures.
However, a substantial gap remains between the promise of
mechanistic toxicology and the actualization of the eld as a
predictive science. For instance, high-throughput in vitro and
in silico toxicity testing remains largely focused on prioritization
of individual chemicals for future investigation allowing to
focus limited resources on the one hand, but which may on the
other hand provide a false sense of safety for “de-prioritized”
chemicals. Specically, these efforts, as well as those aimed at
translating such data into hazard or risk have been hampered
by inadequate coverage of important biological targets given
the limitations of current in vitro methods to simulate in vivo
metabolism or predict effects in different tissues and across
different life stages (Ginsberg 2019), inadequate consideration
of population heterogeneity, and aiming still to provide
assurances of safety rather than quantication of effects across
the population. Furthermore, there has been little progress in
understanding the complex interactions among chemicals and
between chemicals and other intrinsic and extrinsic factors that
affect population health, such as genetics and non-chemical
stressors, including marginalization and other social determinants
of health.
In practice, toxicology largely remains a process based
on reductionist paradigm (Figure 1, left side), classifying
individual chemicals for individual hazards, and investigating
simplistically “linear” mechanistic pathways based on
individual biological targets. Although signicant research and
development investment has been made in improving the
throughput of toxicology through the advent of in vitro and in
silico technologies, the vast majority of these efforts to make
toxicity testing faster, cheaper, and perhaps more relevant are
still fundamentally “one at a time” approaches that feed into
“one at a time” risk assessments and ultimately “one at a time”
decisions. Thus, they ultimately only address a narrow slice of the
human-relevant experiences of toxicity, where 1) all exposures
are time-dependent mixtures of chemical and non-chemical
stressors, 2) every individual has unique susceptibilities and
baseline conditions, and 3) multi-factoral, multi-causal outcomes
are the norm (Figure 1).
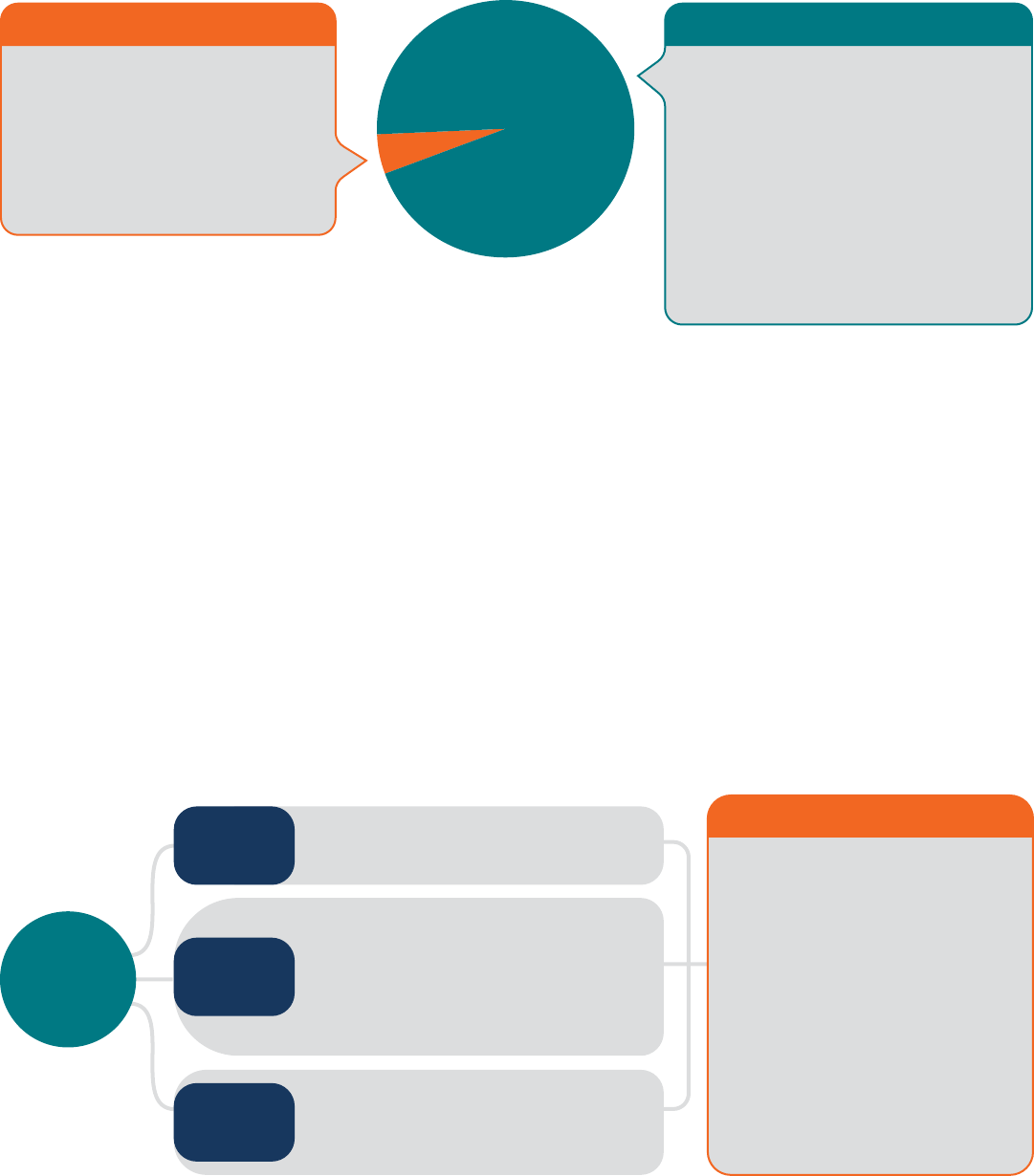
5
This report advocates for a fundamental shift to a holistic
paradigm where toxicology embraces complexity rather than
sweeping it under the rug. Against this backdrop, the workshop
was organized around three main research areas (Figure 2) that
are key to enabling this paradigm shift.
First, whereas both traditional mammalian toxicity testing, and
high-throughput screening assays largely focus on one chemical/
mechanism/outcome at a time, this paradigm shift envisions
toxicology to be exposure-driven, addressing real-life exposure
scenarios in which multiple agents, including social determinants
of health, work together to affect multiple mechanistic pathways
and health outcomes. Additionally, this paradigm shift requires
replacing individual assays that are genetically/epigenetically/
exposomically homogeneous with multiplexed systems that
incorporate inter-cell/tissue interactions on a backdrop of
population variability. Thus, toxicology will become Technology-
enabled, leveraging technological advances from genetics to
bioengineering to enable the characterization of toxicity in
integrated in vitro/in silico platforms across the landscape of
genomics, epigenomics, life-stage, and non-chemical stressors.
Finally, with respect to risk, this paradigm shift requires moving
away from single study-based binary (safe/unsafe) decision-
making to integrating diverse data across multiple data streams
to reach a probabilistic assessment (Maertens, 2022) across
multiple outcomes across the population. Thus, especially with
the emergence of “big data” along with “big compute,” toxicity
will be evidence-integrated, combining multiple evidence
streams across diverse sources of structured and unstructured
information.
The rest of this report summarizes the discussion from the
workshop relating to research challenges, research opportunities,
and the ultimate trajectory to achieve the vision of a holistic,
predictive toxicology.
Figure 1 The proposed paradigm shift in Toxicology research.
Figure 2 The three workshop topics and the expected long-term impacts.
Enabling characterization of toxicity across
genetics, life stage, and non-chemical stressors
and pathobiology of intermediate states,
perturbations, and outcomes while increasing
accuracy, precision, relevance, and domains
of applicability
Frontiers of
Toxicology
Exposure-
driven
Technology-
enabled
Evidence-
integrated
Driven by real-life exposure scenarios and how
multiple agents work together to affect multiple
mechanistic pathways and health outcomes.
Long-Term Impacts
•Safer Chemicals and Drugs through
in vitro/in silico chemical screening,
in vitro/in silico clinical trials,
identification of intrinsic and
extrinsic susceptibilities
•Precision Health through individual,
personalized preventive
interventions, pharmaceutical and
non-pharmaceutical therapies
•Targeted Public Health Interventions
& Environmental Regulations
addressing population & spatial-temporal
variability in genome, epigenome, and
exposome, as well as their socio-economic
consequences
Integrating across diverse sources of structured
and unstructured information with enhanced
access, management, evaluation, and
communication.
Current Reductionist Paradigm
•One chemical at a time
•One endpoint at a time
•One biological target at a time
•Single genetic background tested
•Straight, linear mechanistic pathways
•Interactions simplified or ignored
•Safe vs. unsafe dichotomy
A New Holistic Paradigm
• Directly addressing interactions among
▪ Chemicals
▪ Non-chemical stressors
▪ Heterogeneous populations
▪ Social determinants of health
▪ Life stages, including developmental
origins of disease
•Mechanisms integrated into the complex
physiological networks
•Quantifying probabilistically the impacts on
incidence and severity of human disease
Human-Relevant
Experiences of
Toxicity
A Proposed Paradigm Shift:
Embracing the Multi-Factorial, Multi-Casual Nature of Toxicity
6
Toxicology Research Challenges
For many decades the discussion of changing toxicological
processes was driven by ethical issues of animal use and the
desire to develop and validate so-called “Alternative Methods.”
In the last two decades, it has become increasingly clear that
there are many more reasons to rethink the toolbox of risk
sciences (Hartung, 2017a), namely:
• Long duration and low throughput do not match testing or
public health needs (Hartung and Rovida, 2009; Meigs, 2018)
• Uncertainty in extrapolating results to humans (NASEM, 1983
[the “Red Book”], 1994, 2009 [Science and Decisions])
• Only single chemical/endpoint at a time; does not account
for multiple exposures and non-chemical exposures,
including social determinants (Jerez and Tsatsakis, 2016;
Bopp, 2019; NASEM, 2009 [Science and Decisions])
• Does not account for inter-individual variability (NAS, 2016)
• Does not incorporate associated socio-economic costs and
benets (Chiu, 2017; Meigs, 2018; NASEM, 2009 [Science
and Decisions])
The ongoing transition in terminology in the eld from
“Alternative Methods” to “New Approach Methods” reects
this broader motivation for change. Tox-21c embraced these
challenges and developed a framework of an essentially
mechanistic toxicology of perturbed pathways combined with
quantitative in vitro-to-in vivo extrapolation to human exposure
(Hartung, 2018). A roadmap of consequential steps was
suggested (Hartung, 2009a; Hartung, 2009b).
Workshop participants discussed these overarching challenges
to predictive toxicology and dened the key challenges for
each area as:
Exposure-driven Toxicology
Populations are exposed to multiple environmental agents,
including chemical agents through air, water, food, soil, and
non-chemical agents such as noise, light, and social stressors
(e.g., racism, socioeconomic deprivation, climate). Therefore,
toxicological research that embraces an exposure-driven
approach, characterizing real-life exposure scenarios, including
exposure mixtures and how these agents work together affecting
multiple mechanistic pathways and health outcomes, is needed. A
key opportunity is the expansion of exposomic approaches (Sille,
2020; Huang, 2018; Escher, 2020) to include this broader landscape
of exposures. The workshop participants highlighted three primary
challenges to achieving a more exposure-driven approach:
• Real-world exposures: understanding the interplay
of environmental and social stressors with genetic and
molecular variants
• Predictive intervention: understanding the contributions
of this research toward the identication and evaluation of
effective interventions
• Targeted populations: the inclusion of the affected
communities through participatory research efforts
There are several reasons why an exposure-driven approach
has not yet been embraced. First, many relevant exposures
are not yet fully characterized as we lack the tools and
technologies needed to characterize these exposures, as well
as to understand the health implications. In addition, there has
not yet been a successful engagement of the key stakeholders,
foremost the populations that are directly affected by these
exposures, that is needed for the success of preventive
interventions. However, there are currently substantial advances
coming in these areas and we can easily anticipate substantial
progress in the years to come.
Technology-enabled Toxicology
Predictive toxicology requires expanding the “toolbox” in
several directions. The workshop participants identied the key
challenges to developing the toolbox as:
• Broader model systems: As adverse outcomes involve
interactions of the environment (see above), genes, and
life stage, we need our “model systems” to cover “gene”
and “life stage” more broadly than currently possible using
traditional animal studies (e.g., typically inbred strains) or
even most current high-throughput testing assays (e.g.,
typically based on genetically homogeneous immortalized
cell lines). Example technologies include genetically
diverse population-based in vitro and in vivo resources, and
expansion of experimental designs to cover different stages
of development, as well as developmental origins of health
and disease.
• Access to the intermediate state: Additionally, our
approaches currently cluster at the beginning (e.g.,
high-throughput assays) and the end (e.g., in vivo apical
endpoints) of the pathophysiological process, neglecting
the modulating and stochastic factors that inuence
outcomes that lie between. Thus, approaches that provide
access to intermediate states, perturbations, and outcomes
are needed to better understand the progression to
disease. Example technologies include novel biomarkers,
microphysiological systems (MPS, encompassing organoid
and organ-on-chip technologies), and in silico models (e.g.,
systems toxicology/virtual experiments, AI/ML).
• Assessment tools: We lack the ability to characterize
the predictive accuracy, precision, and relevance of new
approaches or to understand their domains of applicability.
7
Evidence-integrated Toxicology
Toxicology is currently transitioning from a data-poor to a
data-rich science with the curation of legacy databases, “grey”
information on the internet, mining of scientic literature, sensor
technologies, -omics, robotized testing, high-content imaging,
and others. The workshop participants identied the key
challenges to evidence-integrated toxicology as:
• Information sources: There are no established methods
or consensus on how to handle new types of information
sources (which may be incomplete) or how to weigh
evidence strength, risk of bias, quality scoring, etc., or how
to integrate the evidence streams.
• Validation/Verication: In the case of probabilistic risk
assessment, sources of evidence are already integrated,
resulting in a more holistic probability of risk/hazard, so the
challenge is to determine how to validate real-life, t for
purpose, ground-truthing, qualication, and triangulation,
and communicate these probabilities.
• Data Science: We have not yet adopted best practices
for data curation and storage, data mining, analysis, and
visualization.

8
Toxicology Research Advances and Opportunities
The workshop participants anticipate exciting new research
advances on the path to achieving the vision of a holistic,
predictive toxicology that addresses real-life exposure scenarios,
leverages technological advances, and integrates multiple
evidence streams across diverse sources. This section presents
those advances and opportunities according to the three
workshop themes.
Exposure-driven Toxicology
The risks of developing chronic diseases are attributed to
both genetic and environmental factors, e.g., 40% of 560
diseases studied had a genetic component (Lakhani, 2019)
while 70 to 90% of disease risks are probably due to differences
in environments (Rappaport and Smith, 2010). The Human
Genome has been at the center of medical research for the last
forty years, but not many major diseases can be explained or
treated as a result.
Understanding exposure effects and genome x exposure
(GxE) interactions are thus central to the future of medicine.
The original concept of the exposome (Wild, 2016; Vermeulen,
2020), encompassing all exposures of an individual over time,
seems to be impractical and unfeasible as a goal (Figure 3).
The National Academies of Sciences report (NRC, 2010) has
even elaborated on this concept. The 180 million synthesized
chemicals, 350,000 of which are registered for marketing in the
19 most developed countries (Wang, 2020) and myriad natural
and breakdown products seem to make it impossible to measure
and study their effects on humans and the environment. Current
approaches in cells or animals can cost from several thousand to
a million dollars per substance and health effect (Meigs, 2018).
Worldwide toxicity testing covers only a few hundred substances
comprehensively and costs about $20 billion per year. In addition,
human and ecological exposure to substances does not occur
in isolation of single substances or in any constant exposure
scheme. To understand it all or at least a lot of it seems like an
impossible mission.
This has led to a hazard-driven approach to toxicology, i.e., an
established hazard is followed up with exposure considerations
to assess risk. The thresholds of toxicological concern (TTC)
(Hartung, 2017b) concept has been introduced to make
pragmatic use of this by establishing the fth percentile of
lowest-observed (LOEL) or no-observed effect levels (NOEL)
and adding a safety factor of 100. This essentially sets a limit
of possible toxicity at one-hundredth of the point of departure
below 95% of relevant chemicals. For instance, there is a
potential role for TTC to abrogate risk assessment where
exposure and/or bioavailability (internal TTC) (Hartung and
Leist, 2008; Partosch, 2015) are negligible (Wambaugh, 2015),
thus showing a path for how substances could be triaged
according to their negligible exposure. However, TTC-type
approaches are still a “one chemical at a time” paradigm, may
not account for exposures varying temporally or across the
population, and do not address potential interactions among
the thousands of substances to which people are constantly
exposed.
In recent years, the concept of the exposome has been
proposed to capture the diversity and range of environmental
exposures (e.g., inorganic and organic chemicals, dietary
constituents, psychosocial stressors, physical factors), as well
as their corresponding biological responses (Vermeulen,
2020). While measuring those exposures throughout the
lifespan is challenging, technology-enabled advances, such
as high-resolution mass spectrometry, network science, and
numerous other tools provide promise that great advances in
the characterization of the exposome are possible. Indeed,
the exposome approach has achieved traction in recent times
because of the availability of -omics technologies (Sille, 2020)
as discussed below. A NIEHS workshop (Dennis, 2017) saw the
following advantages of an exposome approach:
• Agnostic approaches are encouraged for detection of
emerging exposures of concern
• Techniques, and development of techniques promote
identication of unknown/emerging exposures of concern
• Links exogenous exposures to internal biochemical
perturbations
• Many features can be detected (> 10,000) for the cost of a
single traditional biomonitoring analysis.
• Includes biomolecular reaction products (e.g., protein
adducts, DNA adducts) for which traditional biomonitoring
measurements are often lacking or cumbersome
• Requires a small amount of biological specimen (~100 μL or
less) for full-suite analysis
Figure 3 The exposome concept. [Adapted from Vermeulen, 2020].
9
• Enables detection of “features” that are linked to exposure
or disease for further conrmation
• Encourages techniques to capture short-lived chemicals
• Aims to measure biologically meaningful lifetime exposures,
both exogenous and endogenous, of health relevance
A number of research studies have started to apply these targeted
and untargeted technologies to characterize those complex
exposures and how they impact health and disease, and which
relevant pathways are affected. For instance, birth cohort studies
are attempting to characterize those complex and cumulative
exposures during critical windows that are of increased importance
for long-term human health (Figure 4). Those exposures are not
limited to chemical exposures and consider non-chemical stressors
throughout the lifespan. The concept of cumulative exposures is
critical, as some communities are disproportionally exposed to a
cumulation of chemical and non-chemical exposures which over
time can result in adverse health outcomes.
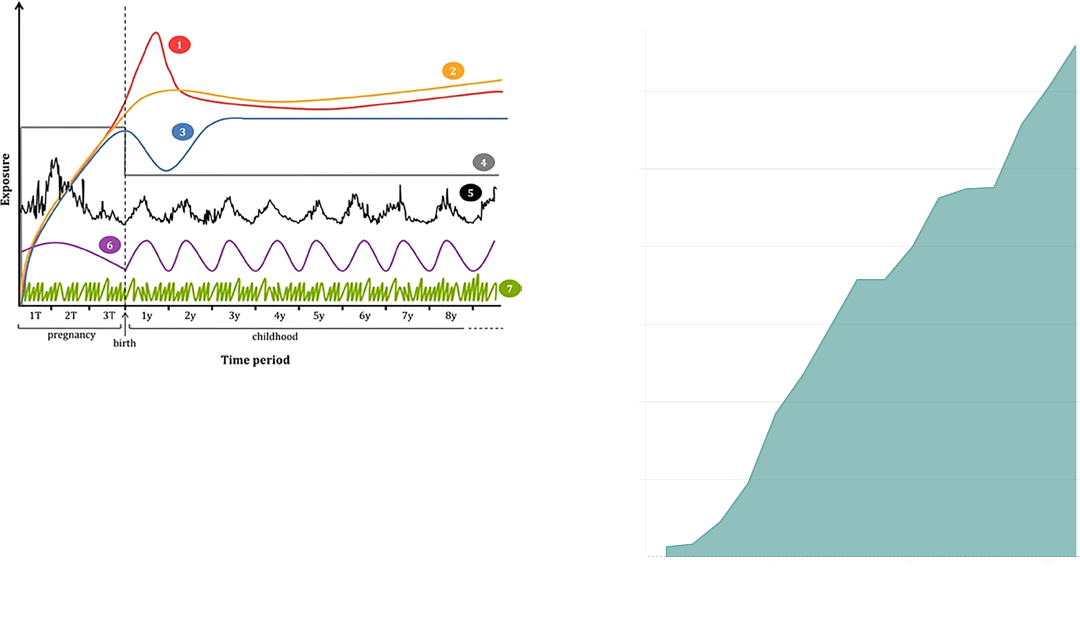
Figure 4 The early life exposome. Examples of relevant exposures and
their exposure patterns during pregnancy and childhood, including
1) persistent organic pollutants (POPs), 2) mercury, lead, 3) arsenic, 4)
secondhand smoke, 5) air pollution, noise, 6) UV radiation, seasonal
exposure to chemicals, 7) non-persistent pollutants. Other exposures
such as psychosocial stressors could follow different exposure patterns.
[Adapted from Robinson, 2015].
These laboratory and exposure sciences advances also require,
in parallel, advancement in biostatistics and data science, to
maximize the information that can be obtained from those
high-dimensional data. For instance, elastic-net regularization
regression is becoming a popular machine learning tool that can
be used to identify the relevant predictors from these complex
sets of exposure data. For instance, these high-dimensional
models were of great relevance to identifying key factors
associated with endogenous intermediate pathways (e.g.,
inammation, protein damage, oxidative stress, and others) in
a pregnancy cohort from Massachusetts called the LIFECODES
cohort (Aung, 2021). These types of cohort studies with complex
exposure data, in diverse populations, prospective follow-up, and
high-quality health outcome data will continue to grow and will
become key tools to advance exposure-driven toxicology.
Technology-enabled Toxicology
The near exponential growth in biotechnology and
bioengineering over the last few decades has created numerous
technologies that could be applied or leveraged in toxicology.
Here we highlight three complementary technology areas that
have the potential to vastly increase the coverage, biological
relevance, and depth of data available for assessing the human
health effects of chemical exposures.
Induced Pluripotent Stem Cell Technologies
The discovery that somatic cells can be reprogrammed
to become pluripotent, recognized by the Nobel Prize in
Physiology or Medicine in 2012, has led to a vast array of
advances in biomedical science, from basic cell biology to
regenerative medicine. Thus, induced pluripotent stem cells
(iPSCs) are among the most substantial research advances of
the 21st century, on par with the sequencing of the human
genome, with thousands of publications per year utilizing this
technology (Figure 5).
Figure 5 Annual growth in publications for “induced pluripotent stem
cells” query in PUBMED, as of July 2022.
In principle, such technologies would enable one to generate
unlimited cells and tissues that retain the genetic information
of the original donor. Cardiomyocytes were one of the rst
functional cell types to be successfully differentiated from iPSCs
and have gone from research lab to commercial application
preclinical safety evaluation of xenobiotics in less than a decade
(Burnett, 2021). They have been found to be useful in identifying
cardiotoxicity hazards for both drugs and environmental
chemicals and are key components of a broad FDA-led initiative
2010 2015 2020
Year of Year
0
500
1000
1500
2000
2500
3000
Count
Annual Pubmed Hits for
"Induced Pluripotent Stem Cells"
10
(CiPA)
3
to address drug-induced arrhythmias, see Figure 6.
However, even for this relatively “mature” technology of
iPSC-derived cardiomyocytes, a number of limitations remain,
including their expressing a more fetal-like phenotype and
challenges in routine and reproducible differentiation from
individual patients. These challenges are even more pronounced
for other cell types, as discussed below.
Nonetheless, the potential for iPSCs to revolutionize biomedical
science overall, and toxicology in particular, is well recognized,
especially when coupled with the rapid development of
advanced in vitro and microphysiological technology.
In vitro and Microphysiological Systems (MPS) Technologies
As discussed in “Toxicology Research Challenges”, there is an
increasing recognition that meeting the needs of toxicology will
require expanding beyond the use of traditional preclinical in
vivo rodent models. These technologies have been termed “New
Approach Methods” (Environmental Protection Agency, European
Chemicals Agency) or "Alternative Methods" (Food and Drug
Administration), and all have the aim of increasing the rigor and
predictivity of toxicity assessments while reducing the reliance
on vertebrate models. Much of the progress in the last 15 years
has been on high-throughput in vitro systems, exemplied by the
Tox21 Consortium
4
, which is a federal collaboration among U.S.
Environmental Protection Agency, National Toxicology Program,
National Center for Advancing Translational Sciences, and the
Food and Drug Administration focusing on “driving the evolution
of Toxicology in the 21st Century by developing methods to
rapidly and efciently evaluate the safety of commercial chemicals,
pesticides, food additives/contaminants, and medical products.”
This effort made use of commercially available assay platforms
across a wide range of targets, testing almost 10,000 compounds.
The screening data generated across a wide diversity of chemicals
and potential mechanisms of toxicity has resulted in hundreds of
publications, with many lessons learned as to the opportunities
and challenges in high-throughput screening data (Richard, 2021).
3 hps://cipaproject.org
4 hps://tox21.gov
5 hps://www.fda.gov/science-research/about-science-research-fda/advancing-alternave-methods-fda
6 hps://mpsworldsummit.com
The workshop participants agreed that more advanced in
vitro technologies, in particular microphysiological systems
(Marx, 2016; Marx, 2020; Roth, 2022), represent the next great
opportunity to advance toxicology (NASEM, 2021). An MPS
model has been dened as one that “uses microscale cell culture
platform for in vitro modeling of functional features of a specic
tissue or organ of human or animal origin by exposing cells
to a microenvironment that mimics the physiological aspects
important for their function or pathophysiological condition.”
5
These may include a wide variety of types of platforms, from
mono-cultures to co-cultures and organoids, and also include
so-called “organ-on-chip” models that include an engineered
physiological micro-environment with functional tissue units
aimed at modeling organ-level responses. These “chip” models
consist of four key components:
• microuidics to deliver target cells, culture uid, waste
discharge
• living cell tissues in either 2D or 3D, including
scaffolding, physical, or chemical signals to simulate the
microenvironment physiologically
• a system for delivering the drug or chemical, either through
the same as the microuidics delivering culture uid, or via a
separate channel (e.g., air-liquid interface)
• a sensing component that may be embedded (e.g.,
electrodes), visual (via transparent materials), or assayed
from efuent
The mushrooming of MPS models has been fueled by stem
cell technologies, 3D cultures (Alepee, 2014), microuidics
(Bhatia and Ingber, 2014), sensor technologies (Clarke, 2021),
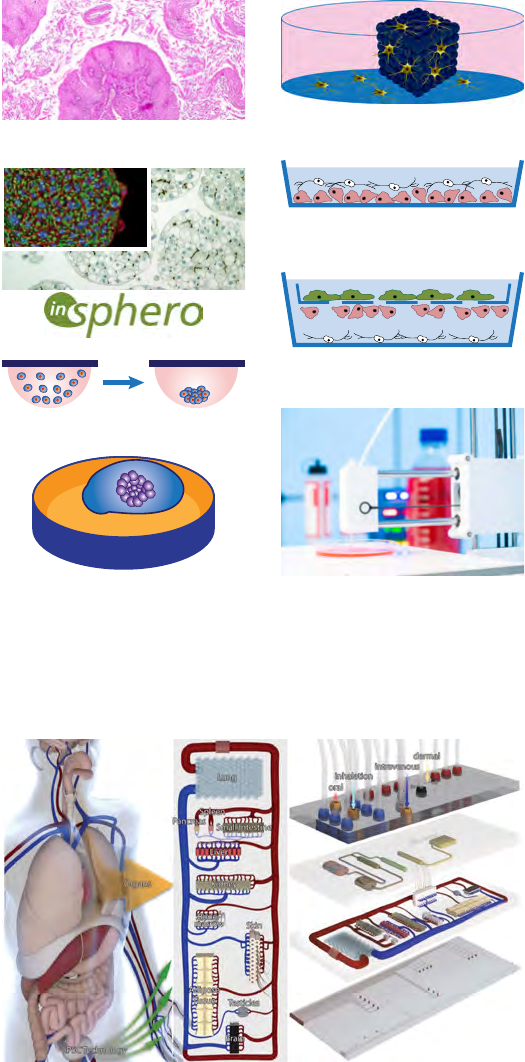
bioprinting (Fetah, 2019) and others. Figure 7 shows different
ways of producing 3D cultures, which are key to creating organ
architecture and functionality as key features of MPS. Notably, the
MPS eld has most recently started to organize itself by annual
global meetings and an International MPS Society.
6
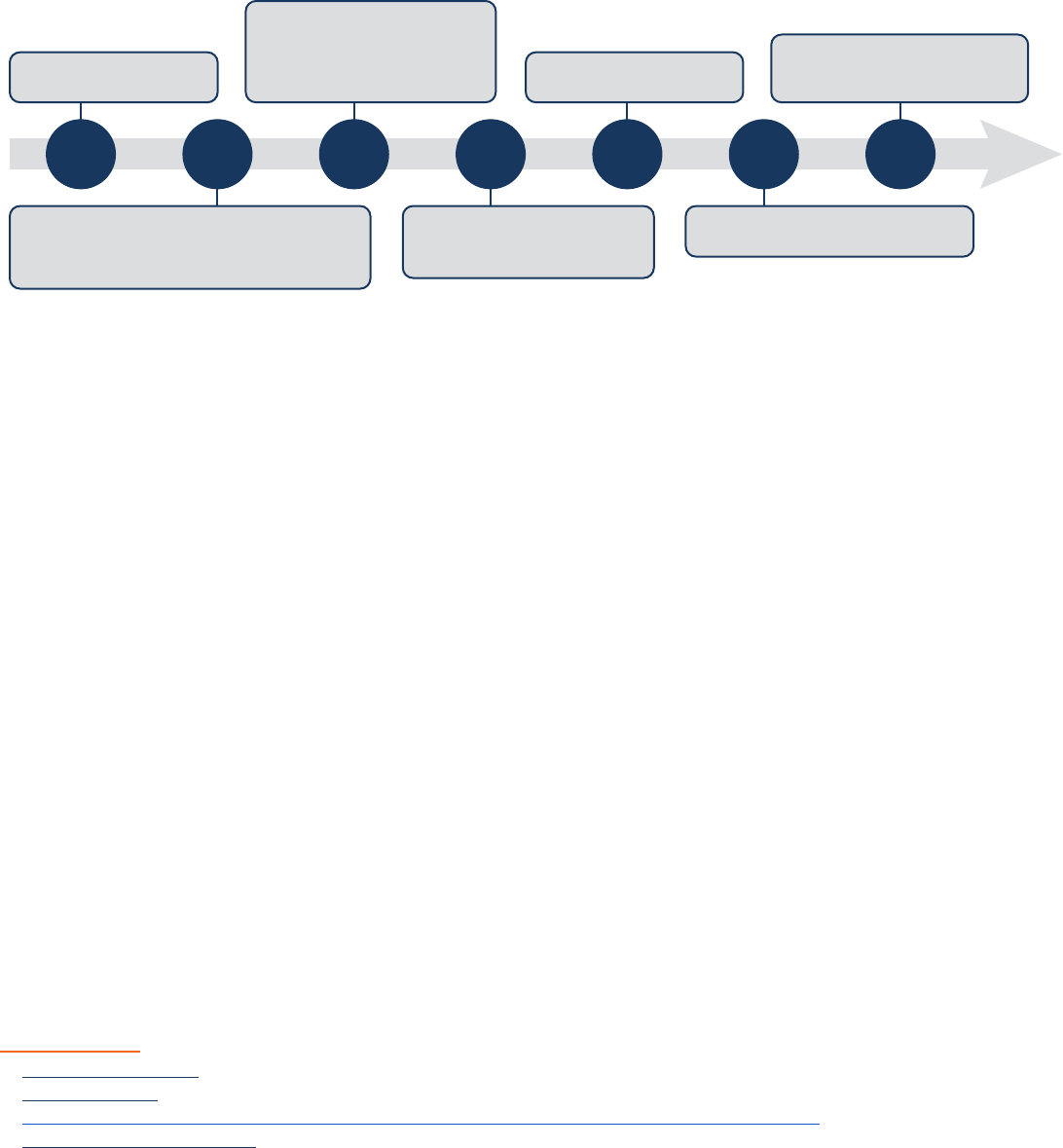
2006 2007 2011 2016 2017 2018 2019
•Generation of iPSCs
from mouse fibroblasts
•Use of patient-specific iPSC-
CMs for disease modeling
•Use of iPSC-CMs for drug
toxicity testing
•Generation of iPSCs from human fibroblasts
•Generation of IPSC-derived
CMs from human fibroblasts
•Use of iPSC-CMs from
patients to recapitulate
clinical toxicity of a drug
•Use of iPSC-CMs for
non-drug toxicity testing
•Use of a healthy population of
iPSC-CMs for drug toxicity testing
•Use of a healthy population of
iPSC-CMs for environmental
chemical toxicity testing
Figure 6 Developmental timeline of induced pluripotent stem cells derived cardiomyocytes (iPSC-CMs) for toxicity testing. [Source: Burnett, 2021]

11
Figure 7 Ways to generate 3D cultures.
MPS models have been developed for nearly every human organ,
and several have been linked together into multi-organ platforms
(Hargrove-Grimes, 2021) (Figure 8).
Figure 8 “Man-on-a-chip.” [Source: Materne, et al., 2013]
Moreover, applications have been reported in drug development,
disease modeling, personalized medicine, and assessment of
environmental toxicants. However, many translational challenges
remain that hinder the application of MPS in toxicology
(Andersen, 2014; Watson, 2017; Nitsche, 2022). Efforts continue
to improve external validation, reproducibility, and quality
control and to enable technology transfer. Noteworthy, Good
Cell and Tissue Culture Practice (GCCP 2.0, Pamies et al., 2022)
has expanded these standards to MPS. Overall, the throughput
remains low, and the cost remains high, hampering broader
application of these technologies, particularly as benchmarking
against simpler in vitro systems has not always revealed
sufcient improvements to warrant the additional time, cost, and
complexity. Nonetheless, emerging efforts to dene appropriate
“context of use” cases for MPS are promising through the
continued interactions among researchers, regulators, and the
private sector (Hargrove-Grimes, 2021; NAS, 2021).
Imaging and Other High-content Measurement Technologies
The high complexity of MPS and consequential lower throughput
make them an ideal match to high-content measurement
technologies, which provide through a comprehensive analysis
of the biological system maximum insight into the Adverse
Outcome Pathway (AOP) in play.
High-content imaging (HCI) combines automated microscopy
with image analysis approaches to simultaneously quantify
multiple phenotypic and/or functional parameters in biological
systems. The technology has become an important tool in the
elds of toxicological sciences and drug discovery because it
can be used for mode-of-action identication, determination
of hazard potency, and the discovery of toxicity targets and
biomarkers (van Vliet, 2014). In contrast to conventional
biochemical endpoints, HCI provides insight into the spatial
distribution and dynamics of responses in biological systems.
This allows the identication of signaling pathways underlying
cell defense, adaptation, toxicity, and death. Therefore, high
content imaging is considered a promising technology to
address the challenges for the Tox-21c approach. Currently, HCI
technologies are frequently applied in academia for mechanistic
toxicity studies and in pharmaceutical industry for the ranking
and selection of lead drug compounds or to identify/conrm
mechanisms underlying effects observed in vivo.
Several ~omics technologies such as genomics, transcriptomics,
proteomics, metabolomics, lipidomics etc. represent further high-
content technologies allowing deep phenotypic characterization
and mechanistic analysis. Hartung and McBride (2011) suggested
earlier the use of combined orthogonal ~omics technologies to
map pathways of toxicity (PoT) (Kleensang, 2014). Noteworthy,
the PoT concept is reminiscent of the AOP approach, which were
both proposed independently in 2011. However, there are some
fundamental differences (Hartung, 2017c): AOP are designed by
experts largely based on their understanding and review of the
literature; they are for this reason very much biased by current
knowledge/belief and typically not quantitative and difcult to
validate experimentally. AOP are narrative, low level of detail,
and largely a linear series of events. PoT, in contrast, are deduced
from experimental data, especially pathway analysis from
untargeted ~omics technologies. PoT are dened on molecular
level with high level of detail, integrating emerging information,
mainly describing network perturbation. They can be studied
further by interventions in the experimental system and often
allow quantitative description. This process is not free of biases
either and the most promising combination of different omics
technologies is still early in development.
Brain Slices
Re-Aggregating Cultures
Colonization of Scaffold
Layered Co-Culture
Hanging Drop
Transwell Culture
Bioprinting
12
Evidence-integrated Toxicology
Toxicology is at the intersection of application and basic science
serving as an integrator of health sciences and public health. While
this is a very powerful position, the dominance of the regulatory
perspective constrains stakeholders. Traditionally, regulation needs
predictions regarding single chemicals, but as a consequence,
risk assessors are stuck in a system where they are tackling one
chemical at a time. Because this is how toxicologists are trained
and how regulatory requirements are formulated, toxicology has
been shaped into a “one-chemical-at-a-time” science. The entire
ecosystem of regulators, the public, and private industry have
ended up focusing on understanding the impacts of each specic
chemical on health or environmental outcomes, even leading
to the creation of trade associations devoted solely to a single
chemical. Breaking out of this paradigm, at minimum, requires
that toxicologists share the data they collect so it can be assessed
and integrated with other data to create a more holistic view of
a chemical’s risk prole. If a risk assessor choses a new tool, its
integration requires broader discussion with regulators and often
regulations must ultimately be updated. Ideally, this discussion and
any information on the tool is public where others can comment
on it. Since there is no centralized effort to do so currently, data
sharing falls to the individual toxicologist and is not necessarily a
common practice. True change has to come from moving public
understanding and regulatory requirements with the eld as one;
which is very difcult to do at the same time.
Big Data
Eighty-four percent of all data in the world has been produced
in the last six years. The scientic literature on the interaction of
humans alone is enormous. For illustration: PubMed is estimated
to cover 25% of biomedical literature. This database includes
about one million new articles per year, of which ~100,000 describe
exposures and ~800,000 include some effects of a substance
on a biological system. Grey literature, such as the internet,
databases of legacy data, -omics technologies, robotized testing,
sensor technologies, image analysis etc. continuously add to
this knowledge base. A critical challenge is in sharing of these
data, which has been a notorious problem in toxicology. Often
information is only in the possession of companies and shared with
regulators in condence, if at all. Not only do we have to overcome
these hurdles, but we also need to establish data collection and/or
meta data standards. This refers in essence to the FAIR principles,
i.e., to make data available in a way that others can use them.
Toxicology is thus currently moving from a data-poor to a data-
rich science, though too many things are still siloed. Raw data
is often behind paywalls or regulatory walls, which can include
being shielded from the public with claims that it contains
condential business information. Consequently, we only see the
tip of the iceberg and data is often not accessible.
Adding to this, no ontologies or metadata allowing people to
make use of each other’s data are available. We generate a lot
of it every day, and generally do not know how to integrate it
unless it is highly curated. Data can be structured by chemical
identity. With more and more data available, the eld of
toxicology becomes dynamic and needs consistent support, e.g.,
to host a central database online. Such a tool needs agreement
regarding how to take data from across datasets. Data needs to
be shareable and usable for machine learning. Ideally, a real-
time assessment would be implemented based on monitoring (for
example integrating data via application programming interfaces
(API)), but such broader integration is hindered by various levels of
technology used by and available in practice. A possible steward,
semantics, standards, and denition of the level of information
needed for human prediction are required. Initially, the focus might
be on narrow chemical spaces with many studies/replicates.
A central problem of safety assessments is how to dene
something as safe. Typically, we have enough data to say
something is toxic, but when do we know enough to say it is
safe? The absence of evidence is no evidence of absence; i.e., a
lack of evident toxicity does not mean that it could not manifest
under different circumstances that are not adequately covered
in the test systems. This calls, on the one hand, for post-
marketing surveillance as done for drugs after market entry, or
more generally for alertness towards consumer feedback and
new scientic ndings.
Systematic Review Methods
A central problem of toxicology is evidence integration (enabling
integration of diverse, cross-disciplinary sources of information)
as more and more methodologies and results, some conicting
and others difcult to compare, are accumulating. This is a
challenge faced in more and more risk assessments, but also in
many systematic review methods that need to combine different
evidence streams (NASEM, 2011, 2017b, 2021; Woodruff and
Sutton, 2014; Samet, 2020; EPA, 2020; EFSA and EBTC, 2018).
Evidence integration is needed on very different levels of data,
studies and to other stressors, as well as across evidence streams.
The central opportunities are in quality assessment and AI,
especially natural language processing (NLP). There needs to
be a common platform, especially on the data side for dynamic
modeling (“dynamic data requires dynamic models”), sharing,
quality control, hardware and software, standards, metadata,
automated annotation, continuous adaptation to AI progress
(e.g. explainable AI), role model evidence-based medicine,
composition of test strategies, validation, and probabilistic risk
assessment. This collaborative open platform to transparently
collect, process, share, and interpret data, information and
knowledge on chemical and non-chemical stressors will enable
real-time and rapid evidence integration, empowering all
steps of protection of human health and the environment.
The combination of tests and other assessment methods
in integrated testing strategies (Hartung, 2013; Tollefsen,
2014; Rovida, 2015), a.k.a. IATA or DA by the Organisation for
Economic Co-operation and Development, needed to integrate
different types of evidence.
Role of Machine Learning and Articial Intelligence
We need evidence integration on the levels of data, information,
knowledge, and ultimately action. A system for integrating across
different levels of information that are each integrated within

13
their own space, requires broad integration of quality information
with the proper infrastructure to support this. The vision is to
create an infrastructure with harmonized agreement on the
levels of information and for what they may be best suited. For
this and its broad use, more toxicologists with computational
skills are needed. We also need common vocabularies across
different levels of information, data architectural standards for
release and utilization, real-time integration (e.g., through APIs),
and annotation at different levels. Such annotation requires
the connection of raw data to study metadata and the use of
language that a computer can digest by NLP through ontologies,
standardized “controlled” vocabularies, harmonized templates
such as IUCLID (https://iuclid6.echa.europa.eu/), and a library of
synonyms. A major question is what can be done to make sure
data is encoded/tagged to make it useful? High-quality training
sets for annotation to build knowledge graphs, causal networks,
etc. need to be developed. However, the use of annotation/
structured databases is an old way of looking at things. It is
almost impossible to get people to conform to data annotation
guidelines, so instead the eld will need to embrace methods to
manage unstructured data.
We might rather spend energy on building better NLP and deep
learning technologies to analyze unstructured data. The enormous
progress on NLP in recent years means that we are moving very
close to surpassing the Turing Test, if we have not already. The
Turing Test is a deceptively simple method of determining whether
a machine can demonstrate human intelligence. If a machine can
engage in a conversation with a human without being detected as
a machine, it has demonstrated human intelligence. Over the last
two years, enormously large models have been trained (Hoffmann,
2022). They use 140 to 530 billion parameters and 170 billion to 1.4
trillion training tokens. Some of these models claim to have been
trained on the entire Internet. They can respond in real-time to
questions with high accuracy, write articles indistinguishable from
those by human authors and even write code for computers. The
rst impact of the NLP breakthrough is that human knowledge
becomes machine-readable. Our vision is that this enables the
creation of similar models to virtually grasp the interaction of
organisms with chemical substances.
Toxicologists can read and extract information better, but
a computer can do this faster on many more sources. We
now need to train computers to be as good as humans in
interpreting data. AI is the best tool for evidence integration,
and evidence integration must become the standard for risk
and safety assessments. The big question is: how are people
going to use this information generated by AI? Here we need
to separate our vision from its implementation. Toxicological
research areas and associated S&T advances can overcome
hurdles to enable toxicology as a predictive science via evidence
integration. From the explosion in the use of machine Learning
and data science, the emerging use of NLP, knowledge graphs,
and next-generation-omics analytics we need to move to
explainable AI, embrace reinforcement learning and modern
database management. The platform to be established will
7 hps://www.ebtox.org
need an IT architecture, hardware and software, continuous
deployment/support, decision support tools, expert systems etc.
Evidence-based methodologies as furthered by evidence-based
toxicology
7
(e.g., systematic review principles, risk of bias, meta-
analysis, quality scoring, probabilistic approaches) can serve as
role models for objective and transparent handling of evidence.
Besides making sense of evidence pieces, such a platform can
also guide the composition and validation of Testing Strategies
(IATAs, DAs, AOP networks) and extraction of human relevant
reference datasets.
Probabilistic Approaches
Recognizing that as science delivers only probability rather
than absolutes, probabilistic tools lend themselves to all of
these (Maertens, 2022; Chiu and Paoli, 2020), will enable us to
move away from black/white, toxic/non-toxic dichotomies, as
well as better support life cycle and socioeconomic analyses
that require evaluation of incremental benets or risks rather
than “bright line” evaluations (NASEM, 2009; Chiu, 2017;
Fantke, 2018, 2021). Substantial progress on developing and
implementing probabilistic risk assessment approaches has
been made in the last 10 years (Chiu and Slob, 2015; Chiu,
2018), with the publication of guidance from the WHO/IPCS
(World Health Organization & International Programme on
Chemical Safety, 2018). Conceptually, this involves replacing
the xed values currently used for both the initial 'point
of departure” dose, as well as the “uncertainty factors”
with distributions that reect the state of the scientic
understanding, incorporating and combining uncertainties
quantitatively through statistical approaches (see Figure 9
for example applied to the Reference Dose). Several case
studies illustrating the broad application of probabilistic
approaches have been demonstrated (Blessinger, 2020; Chiu,
2018; Kvasnicka, 2019). Moreover, this conceptual approach
to deriving toxicity values probabilistically can be extended
to non-animal studies (Chiu and Paoli, 2020), as well as to
incorporating population variability through genetically diverse
models described above (Chiu and Rusyn, 2018; Rusyn, 2022).
In this way, probabilistic approaches provide a framework that
facilitates integration across different data types and sources.
14
Point of
Departure
Divide
by 10
Divide
by 10
Reference
Dose
Traditional Approach
POD
POD
10
POD
100
RfD =
Test System
(e.g., experimental animal, in vitro assay)
Inter-Species or
IVIVE Adjustment
Intra-Species
Variability
“Typical” Member of
Human Population
Common Conceptual Model
“Sensitive” Member of
Human Population
Dose or Concentration Distribution with
Effect Size M in Test System
Inter-Species or
IVIVE Distribution
Median to I
th
Percentile
Distribution
Dose Distribution with
Effect Size M in Median Human
Probabilistic Approach
Dose Distribution with
Effect Size M in I
th
Percentile Human
Reference Dose (RfD):
An estimate (with uncertainty spanning perhaps an order of
magnitude) of a daily oral exposure to the
human population
(Including sensitive subgroups
) that is likely to be without
an appreciable risk
of deleterious effects during a lifetime.
Probabilistic RfD (PrRfD):
A statistical lower confidence limit on the human dose
that at which a fraction I of the population shows an effect
of magnitude (or severity) M or greater (for the critical
effect considered).
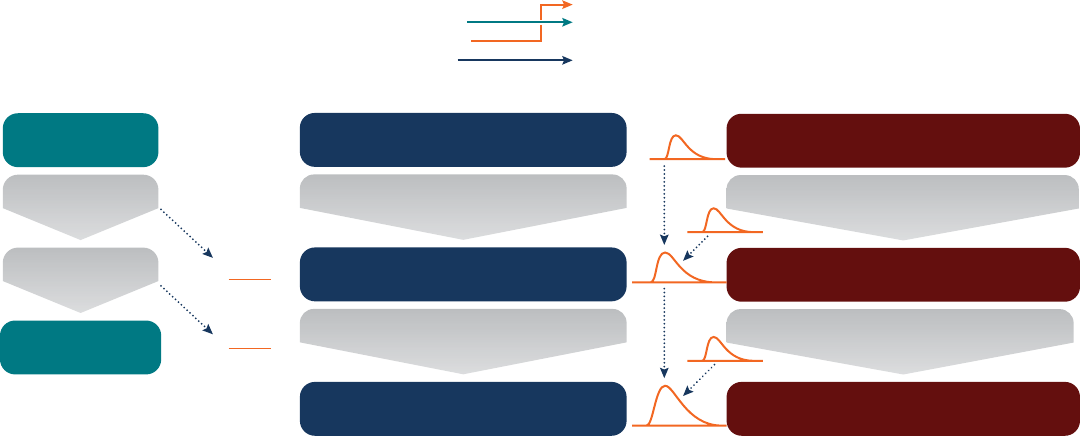
Figure 9 Illustration of the transition from deterministic to probabilistic approaches when deriving reference doses from toxicity data. The
“Traditional Approach” refers to the practice attributed to Lehman and Fitzhugh (1954) to derive a "safe dose” by taking the dose level without
signicant effects in an animal study (a “point of departure”) and dividing by a “safety factor” of 100. The “Common Conceptual Model” is an
abstraction of this this procedure, whereby information from a test system (whether animal study or other type of data) is rst adjusted to the
“typical” in vivo human, and then adjusted to account for human variability in susceptibility, thereby deriving dose level that is protective of
“sensitive” members of the human population. The “Probabilistic Approach” further incorporates quantitative uncertainty and variability into this
conceptual model, using probability distributions at each step instead of single numbers, so that the result is a distribution (reecting incomplete
knowledge) for the dose that would cause on effect of magnitude “M” in the “I”th most sensitive percentile of the human population. [Adapted
from World Health Organization & International Programme on Chemical Safety., 2018]
15
Toxicology Research Trajectory
Recognizing the broad research advances and opportunities that
have arisen in the last 15 years since the 2007 Tox-21c report,
the workshop participants outlined their vision for the future
research trajectory needed to fulll the promise of transforming
toxicology into an exposure-driven, technology-enabled,
evidence-integrated eld that can better address population and
precision health while ensuring safe pharmaceuticals and a safer
environment. For each of the three research areas, participants
delineated a 5-, 10-, and 20-year plan for building capabilities
that would facilitate this transformation.
Exposure-driven Toxicology
Workshop participants identied several major areas of research
focus to advance exposure driven-toxicology in the coming
decades: 1) real-world-based exposure designs, 2) population-
scale measurements, 3) strategies to ask the right questions, and
4) consideration of ethical and policy implications.
Real-world-based Exposure Designs
Developments in this area are needed to allow for better in
silico, cellular, organoids, model organisms, as well as full
populations-based longitudinal studies. These developments will
allow studies to be conducted in a way that supports prediction
of environmental transport and fate (including chemical
transformations, inter-species comparisons, the application of
the understanding of exposure levels and exposure mixtures)
relevant to the population and its sub-groups. They will also
allow us to apply that knowledge to the experimental setting.
By using this real-world-based exposure design, the results
of different approaches to answering similar questions will be
easier to compare, and make it easier to utilize triangulation as a
key strategy for assessing the health effect and relevant toxicity
pathways of chemical and non-chemical exposures.
Population-scale Measurements
To understand the relevant exposures that lead to disease in
general and specic populations, additional efforts are needed to
develop biobanks (including biological specimens) and ecobanks
(including environmental samples) that inform on the distribution
of thousands of chemicals and non-chemical stressors in relevant
populations. Factors of interest include relevant exposure
scenarios, sociodemographic conditions, and relevant disease or
health status. Beyond human populations, the inclusion of animals
and the ecosystem for real-world exposure assessment is of
relevance to human environmental health, as well as environmental
health and toxicology, more broadly. Recent studies, for instance,
have shown that exposure assessment efforts in companion
animals, such as cats using non-invasive silicon tags, can contribute
to the assessment of ame retardants in homes, and their potential
role in feline hyperthyroidism (Poutasse, 2019).
Ask the Right Questions
One of the complexities in the current eld of omics technology
is how to prioritize the right questions in a way that leads
to the correct computational approach. For instance, the
question might be related to the total mixture, or to specic
components of a mixture. Thinking strategically and with the
right stakeholders (community, policymakers, interdisciplinary
scientists), will contribute to developing those right questions
in ways that are most useful for society and respecting
privacy concerns that many have regarding the unintended
consequences of data sharing.
Ethical and Policy Implications
An important amount of the workshop discussion focused
on aspects related to the ethical and policy implications of
toxicological research including the disproportionate burden
of exposures affecting disadvantaged communities. Groups
discussed the need for research to address those concerns by
incorporating elements of community engagement, citizen
support and environmental justice that must keep pace with
the technology.
Anticipated Capabilities
Regarding the key anticipated capabilities for exposure-driven
toxicology, the workshop participants anticipated the following
achievements as shown in Table 1 and described here:
At 5 years:
• Scale-up technology for high quality inexpensive assessment
of 1000-5000 chemicals that can be tagged to exogenous
exposures including non-chemical stressors. Technology is
currently slow and throughput is not high enough, which
makes exposomic approaches expensive.
• Develop libraries that tag key information for those chemicals
(meta data layering) to ensure their interpretation. There is
currently a lack of validation for many chemical signatures that
can be identied with untargeted technologies as to their
prediction of health effects.
• These technology problems can be solved through effort
and investment, similar to the genome project.
At 10 years:
• The availability of scalable technology for exposomics to
achieve high throughput, that is also cheap, sensitive, and
specic will allow us to apply this exposure-based approach
to longitudinal studies and biobanks.
• Studies that can be both retrospective and prospective
ensuring “FAIR” ness and linking exposome with health
outcomes.
• Retrospective studies will allow us to go back decades and
leverage biobanks. At the same time, we will be able to plan new
prospective studies to evaluate the exposures of the future.
20 years:
• Exposome-disease prediction will integrate detailed exposure-
based information with health outcome data in large scale
and numerous populations. We will achieve a great level of
precision in disease prediction that will be environment-based
and can also leverage gene-environment interactions.
• This knowledge will provide us with new forms of exposome
targeted prevention and treatment.

16
Table 1: Timeline for Key Exposure-driven Toxicology Developments
Key Capability Near-term (5-yr) goal Mid-term (10-yr) goal Long-term (20-yr) goal
Analytical
chemistry
Exposome assays (1000—
5000k/person)
High throughput exposome
assays (10,000/person)
Exposome disease prediction
Metabolomics,
toxicology
Reference exposome library
(meta-data layering)
Organ specic disease associated
Exposome targeted
treatment and prevention
Epidemiology,
clinical research
Disease associated metabolites
Retrospective and prospective
studies ensuring “FAIR” and linking
exposome with health outcomes
Exposome targeted treatment
Technology-enabled Toxicology
Workshop participants anticipate new technological capabilities
in two key research areas to fulll the promise of transforming
toxicology (see Table 2). These include biological capabilities
to provide a diversity of cells and tissues and bioengineering
capabilities to develop relevant and reproducible assays.
Moreover, in each, a set of supporting computational capabilities
will need to be developed.
Biological Capabilities
The critical path for biological capabilities lies in the
understanding of heterogeneity and susceptibility throughout
the life course at multiple scales from cells to the whole
organism. As genetics have turned out to be a much smaller
factor in outcomes than originally anticipated, there is a
need to better understand how non-genetic factors, such
as epigenetic differences and social and environmental
stressors, individually and collectively modulate development,
pathology, and pathophysiology. Because of the diversity in
the human population, an important resource for enabling this
understanding will be reliable and reproducible sources of cells
from multiple tissues representative of the population. Cell
sourcing is particularly important because it is likely that in vitro
microphysiological and other bioengineered systems (discussed
below) will play an essential role in untangling these complex
interactions. It is also recognized that in parallel, computational
capabilities, including multi-omic databases and advances in
interpretable AI, will need to be developed to move biological
capabilities forward.
Bioengineering Capabilities
With respect to bioengineering capabilities, the main
hurdles are: the lack of validated and standardized platforms
with automated fabrication, and the lack of availability of
individualized cell differentiation to enable personalized
toxicological evaluation. It is recognized that a range of t-
for-purpose models, ranging from simple suspensions and
monolayers to fully vascularized and innervated multi-organ
microphysiological systems, will be developed over time.
However, for any of them to be personalized, cell differentiation
protocols are needed that can enable creation of multiple
tissues from iPSCs from any individual. Coupled with the
biological capabilities understanding heterogeneity and
susceptibility, these bioengineering capabilities would enable
modeling of the diversity of the human population through
time. If the goals of automated fabrication and low cost are
also achieved, then a “multiverse”-type model platform is
envisioned, in which each person could have numerous “chip-
based twins” that could predict a range of possible future states
depending on different future exposures. As with biological
capabilities, a parallel set of computational capabilities will
be required, with the goal of creating “digital twins” to go
alongside the suite of “chip-based twins.”
Table 2: Timeline for Key Technology-enabled Toxicology Developments
Key Capability Near-term (5-yr) goal Mid-term (10-yr) goal Long-term (20-yr) goal
Biological
Representative and reliable
sources of human cells
National repository of human
cells representing key tissues
International repository of
human cells representing
all major tissues
Repository of human cell
types that is representative
of population diversity
Determinants of
heterogeneity and
susceptibility to toxicants
Role of genetic, epigenetic,
and social determinants
Understanding impact
of timing of exposure
and life stage
In vitro/in silico models for
heterogeneity and susceptibility
Understanding of
pathology, pathophysiology,
and development at
multiple scales
Faithful in vitro cell
differentiation coupled with
integrated multi-organ systems
Non-invasive, label-
free biomarkers
Integrated biomarkers
for modeling normal and
diseased organismoids.
Computational support for
biological capabilities
Multi-omic databases that are
comparable and computable
Integrated multi-omic databases coupled with interpretable AI

17
Evidence-integrated Toxicology
Evidence integration (animal studies, human studies, in vitro,
and all other types of studies) is needed at different levels of
integration, including: raw data, reports/scientic papers (meta-
analysis), and data on exposure and hazard. We integrate raw
data by interpreting it, and transforming it into information. We
integrate different pieces of information in our studies and reports
and create knowledge from the body of available studies and
papers. Ultimately, we act based on this knowledge, but where
do we integrate? Integration across evidence levels requires a
systematic review, based on structured data submissions. Curated
data sources and models fall between raw data and study reports.
The hope is that twenty years from now we will have a system
for integrating all these data and factors. This would enable
better risk management for public health with enormous societal
consequences. But we will also need to address the challenges of
communicating the results. Any evidence integration that does not
lead to simple classication is a potential communication problem.
However, chemicals cannot be simply put into black and white
bins (toxic vs. non-toxic) because they exist on a spectrum best
characterized by the probability of hazard.
To make Big Data common and encourage its use broadly
in toxicology in 10-20 years, we must incentivize people to
share information that today is still considered proprietary to
businesses. A lot of companies are already required to submit
structured information as regulatory data, but there needs to be
regulatory change to make the information accessible (possibly
with acceptable “blinding” of certain data). A key role of public-
private partnerships for data sharing, accessibility, modeling,
and cross-sector harmonization was identied. The application
of blockchain technology to encrypt pharma/health data and
federated model building should be explored. For the animal
part of toxicology, we have a moral obligation to make results
public. Experimenters received an exemption from society to
do something they normally would not (harm an animal), so
they owe society the data/outcome in return. The OECD QSAR
Toolbox is a pioneering tool that could serve as a model since
industry and regulators are less often using tools offered by
vendors, especially those tools which use commercial data that’s
not publicly available. In addition, there are differing levels
of transparency requirements (open access vs. open data vs.
open source). Cost could also be an issue, and promoting data
collection, processes, and trusted data brokers will be needed.
Training models with synthetic data sets as done more often with
patient/clinical data might be a promising avenue to explore.
The FAIR principles (Wilkinson et al., 2016) give important
guidance for the accessibility of structured and annotated data
to make them useful. Efcient utilization relies on real-time data
integration and updates as well as structured, annotated input
data to unlock the information contained.
The opportunities come from Big Data (characterized by the
3V's, or volume, variety, and velocity). It is expected that NLP for
data extraction will play a big role. Currently, most AI is a black
box. To make progress, we will need to move from this paradigm
to an explainable AI. Increasingly there is a need to combine
prospective and retrospective testing strategies. Policy does not
keep pace with the speed of technology. Strategies for enabling
evidence integration, such as better integration of various
~omics data from mechanistic toxicology, ~omics data layering
at the individual level is needed. We need to understand how to
incorporate less understood but equally important variables into
the equation, such as microbiome, circadian rhythm, and age. A
goal is to develop mechanistic models by changing from data-
driven to mechanism-driven. Challenges include:
• Finding incentives and reward structures in institutions to
encourage integration of big data approaches
• Dening relevant toxicants information to move through
three areas of measurement: time, space, and population.
This includes expansion to global populations.
• Small portable technologies to capture genetic and
environmental heterogeneity
Table 2: Timeline for Key Technology-enabled Toxicology Developments
Key Capability Near-term (5-yr) goal Mid-term (10-yr) goal Long-term (20-yr) goal
Bioengineering
Individual-specic cell
differentiation protocols
Reproducible cell differentiation
for all essential tissues for
at least one donor
Multi-donor cell differentiation
for all essential tissues
Individualized cell differentiation
for all essential tissues
Menu of t-for-purpose in
vitro model platforms.
Models representing
population diversity
Models representing
both health and
diseased populations
Models capable of
modeling “multiverse” of
potential future states
Validated, reproducible,
and affordable multi-organ
microphysiological “chips”
Platform standardization and
automated fabrication for a
dozen commercial platforms,
validated for individual organs
Commercial/off-the-
shelf single-organ chips;
validated multi-organ
chips; demonstrated
personalize chips
Validated personalized
“multiverse” chip
Computational support
to bioengineering
Quantitative in vitro to in
vivo extrapolation coupled
with in silico modeling
of model platforms
Digital/in silico
organ modeling
Personalized digital twins

18
• Designing a resource-efcient tiered strategy composed of
various methods for gathering human data for supervised
learning
• Identication of enabler technologies that make analyses
affordable and reliable
• Integration of human, animal, in vitro, etc. studies in
cohesive projects, allowing crosstalk and feedback
• Testing, data analysis, treatment and prevention informed by
predictive modeling (recursively applied to testing)
• Cloud-based computing platforms for continuous data and
model integration
• Common language so that information can move between
the three areas of exposure, technology, and evidence-
integration
• Robust informatic infrastructures: graph databases and novel
ML over structured and non- structured
• Need for Quality Assurance/Quality Control and validation
• Quality control of MPS reporting input into AI
Table 3 Timeline for Key Evidence-integrated Toxicology Developments
Key Capability Near-term (5-yr) goal Mid-term (10-yr) goal Long-term (20-yr) goal
CompTox literacy
Trainers and trainees
with CompTox skills
Establish curricula broadly
Highly skilled CompTox
workforce
NLP for decision
making
Annotation enabling resources
(e.g., synonyms, ontologies,
standardized vocabularies); NLP
to automatically retrieve/parse
study methods [unstructured data];
NLP to learn about relationships
in currently structured data
NLP to automatically
extract/parse study results;
accessible networks of NLP-
dened causal networks
NLP to automatically interpret/
combine study results; NLP
associations create datasets
that inform risk assessments
Data sharing
Global use of IUCLID and other
structured repositories and APIs
Platform of networked annotated
databases; community
engagement in exposure/health
data sharing and tracking
Real-time update and analysis
via networked platform
Explainable AI for
evidence integration
Explainable AI algorithms
and modeling pipelines
Interactive decision-support tool
that integrate evidence streams
Widespread implementation
of AI/ML in decision making;
probabilistic risk assessment

19
Accelerating Progress
8 hps://marianamazzucato.com/books/the-entrepreneurial-state
9 hps://web.ornl.gov/sci/techresources/Human_Genome/project/budget.shtml
10 hps://web.ornl.gov/sci/techresources/Human_Genome/project/economics.shtml
Realizing the Tox-21c 2.0 vision needs the Entrepreneurial State
8
(i.e., massive government investment) to create dedicated grants
for research, training, and implementation, in addition to efforts
to overcome institutional barriers. The evident role model is the
Human Genome Project, which largely transformed biomedical
science (Hood, 2013). Its costs have been estimated at up to
$3 billion.
9
A Battelle report from 2011 appraised the large and
widespread economic and functional impacts
10
: “Between 1988
and 2010 the human genome sequencing projects, associated
research and industry activity—directly and indirectly—generated
an economic (output) impact of $796 billion, personal income
exceeding $244 billion, and 3.8 million job-years of employment.
In the 2013 update, these numbers increased to economic
(output) impact of $965 billion, personal income exceeding
$293 billion, and 4.3 million job-years of employment.” The
transformation of toxicology envisioned here as a Human
Exposome Project (HEP) represents a similar opportunity,
promising to identify an even larger fraction of causes of disease
and opening up new opportunities for prevention and cure.
Governance: The necessarily multi-disciplinary and international
character of toxicology requires strategic steering. A cross-
agency alliance in the US could play a central role. New metrics
for success that measure impact/implementation must be
developed. Public-private partnerships are the most promising
avenue given their economic prospects. In its build-up,
dedicated preparatory programs, infrastructure, partnerships,
and engagement with the community have to be developed.
To start, a centralized effort is needed to facilitate data sharing
and to create the analysis platforms discussed in the evidence
integration section.
Education and Training: The challenge in educating and
training skill set development is a cross-cutting issue. The lack
of multidisciplinary skills was mentioned across all workshop
groups, as was the need to strike a balance between what you
can and cannot share (privacy, IP, crowdsourcing). A continued
dialogue through workshops, papers, strategic plans, policy
advice, and implementation is needed. The implementation
especially requires training in computational toxicology skills and
user comfort with computational tools. Most university curricula
are not up-to-date in this respect. The younger generation
is very receptive. A portfolio of CompTox training materials
would be helpful. It is also a communication challenge to both
the workforce and through outreach/engagement (two-way to
understand needs; multi-directional communication and training
on all levels is key) to the public and decision-makers.
Communication: The assembly of the components for
transforming toxicology and revamping risk sciences is a premier
communication challenge. Already, personal risk and public
health are difcult to communicate to the general public and
policymakers. Identifying drivers of disease must present itself
not as an anti-industry stance, but a societal need with business
opportunities. Communication with end-users to understand and
overcome institutional barriers is required. We need to articulate
in terms of the problems we are trying to solve.
20
Conclusion
This workshop is visionary, looking 10-20 years into the future.
Incredibly powerful novel methodologies exist to revamp
toxicology; the challenge is their implementation. We have to
enable toxicology to keep pace and benet from cross-sector
advances. There is a need for proof-of-concept examples
around information retrieval, evidence integration, quality
assessment, and decision support. A special opportunity lies in
crowdsourcing. Investment in technical infrastructure will facilitate
decision support tools (interpretable, actionable, probabilistic,
exible) that integrate multiple evidence streams.
The implementation of this vision is based on several key
expectations:
1. Biomonitoring and exposomics can evolve and be scaled to
make toxicology and environmental health more exposure-
driven
2. Relevant human model systems can be bioengineered to
study disease etiologies and interventions, especially of
exposure, as microphysiological systems
3. Computational approaches allow us to scale assessments of
chemicals and drugs
4. Evidence integration from these disruptive technologies can
guide risk assessment and management
Long-term Impacts
Ultimately advances in these areas would enable transformation
in toxicology with lasting impacts in three major ways:
• Safer Chemicals and Drugs. Much of toxicology today is
focused on ensuring the safety of xenobiotic exposures,
whether they be pharmaceuticals intentionally administered
or chemicals to which one is incidentally exposed through
the environment, consumer products, or occupation. Thus,
the most direct impacts of predictive toxicology would be
improvements in safer chemicals and drugs through higher
throughput and/or higher relevance in vitro or in silico
assays, particularly ones that are better at identifying intrinsic
and extrinsic susceptibilities.
• Precision Health. Similarly, there is substantial investment
already in precision medicine in the form of personalized
drug treatment. However, the research described here could
broaden this to the concept of precision health, which would
not only include personalized pharmacological treatment,
but also personalized preventive interventions and non-
pharmaceutical therapies. Moreover, while current efforts in
precision medicine focus on pharmacogenomics or poly-
pharmacy, advances in the frontiers of toxicology discussed
above could enable that individualization to extend to
the epigenome and exposome, as well as to interventions
connected with community health and well-being including
access to green-spaces, clean air and water.
• Targeted Public Health Interventions and Environmental
Regulations. Finally, the concepts of precision health
could be expanded to support public health with better
targeted public health interventions and environmental
regulations. This would better elucidate the toxicological
impacts of the genome, epigenome, and exposome,
all in combination. Thus, not only could there be
better assurance that interventions and regulations are
protective of the most vulnerable, but novel approaches
may also be revealed that enable better targeting of such
measures so that scarce resources can be allocated to
achieve the greatest overall benet.
Ultimately, the workshop participants envision a future for
toxicology as a Human Exposome Project in which collaborative,
technology-enabled open platforms transparently generate,
collect, process, share, and interpret data, information, and
knowledge of real-world chemical and non-chemical stressors to
enable real-time and rapid evidence integration, empowering all
steps of protection of human health and the environment.
21
Glossary
This is the section provides the denitions for terms used in the body of the report (see also Ferrario et al.,2014; Sille et al., 2020).
Adverse Outcome Pathway (AOP): An AOP is a sequence of events from the exposure of an individual or population to a chemical
substance through a nal adverse (toxic) effect at the individual level (for human health) or population level (for ecotoxicological
endpoints). The key events in an AOP should be denable and make sense from a physiological and biochemical perspective. AOPs
incorporate the toxicity pathway and mode of action for an adverse effect. AOPs may be related to other mechanisms and pathways as
well as to detoxication routes.
Biomarker: Indicator signaling an event or condition in a biological system or sample and giving a measure of exposure, effect, or
susceptibility.
Biomonitoring: The measurement of the body burden of toxic chemical compounds, elements, or their metabolites, in biological
substances
Evidence-based toxicology (EBT): EBT is a process for transparently, consistently, and objectively assessing available scientic
evidence in order to answer questions in toxicology. Particularly EBT: a) promotes the consistent use of transparent and systematic
processes to reach robust conclusions and sound judgments; b) displays a willingness to check the assumptions upon which current
toxicological practice is based to facilitate continuous improvement; c) recognizes the need to provide for the effective training and
development of professional toxicologists; d) acknowledges a requirement for new and improved tools for critical evaluation and
quantitative integration of scientic evidence; e) embraces all aspects of toxicological practice, and all types of evidence of which use
is made in hazard identication, risk assessment, and retrospective analyses of causation; f) ensures the generation and use of best
scientic evidence; g) includes all branches of toxicological science: human health assessment, environmental and ecotoxicology, and
clinical toxicology; h) has the potential to address concerns in the toxicological community about the limitations of current approaches
to assessing the state of the science; i) acknowledges and builds upon the achievements and contributions of Evidence Based
Medicine/Evidence Based Health Care.
Exposome: Concept describing the totality of exposure experienced by an individual during their life and the health impact of those
exposures (Wild, 2005), redened (Miller and Jones, 2014): The cumulative measure of environmental inuences and associated
biological responses throughout the lifespan, including exposures from the environment, diet, behavior, and endogenous processes.
Hazard: 1) A biological, chemical, or physical agent with the potential to cause an adverse health effect (European Commission, 2002).
2) The inherent characteristic of a material, condition, or activity that has the potential to cause adverse effects to people, property, or
the environment.
Metabolomics/Metabonomics: Evaluation of cells, tissues, or biological uids for changes in metabolite levels that follow exposure
to a given substance in order to determine the metabolic processes involved, to evaluate the disruption in intermediary metabolic
processes that results from exposure to that substance, or to determine the part of the genome that is responsible for the changes.
Note: Although “metabolomics” and “metabonomics” are frequently used as synonyms, there is a growing consensus that there is a
difference in that “metabolomics” places a greater emphasis on comprehensive metabolic proling, while “metabonomics” is used to
describe multiple (but not necessarily comprehensive) metabolic changes caused by a biological perturbation.
Risk assessment: A scientically based process consisting of four steps: hazard identication, hazard characterization, exposure
assessment, and risk characterization.
Threshold of toxicological concern (TTC): Human exposure threshold value for a group of chemicals below which there should be no
appreciable risk to human health.
Toxicokinetics: Generally, the overall process of the absorption (uptake) of potentially toxic substances by the body, the distribution
of the substances and their metabolites in tissues and organs, their metabolism (biotransformation), and the elimination of the
substances and their metabolites from the body. In validating a toxicological study, the collection of toxicokinetic data, either as
an integral component in the conduct of non-clinical toxicity studies or in specially designed supportive studies, in order to assess
systemic exposure.
Validation: The process by which the reliability and relevance of a particular approach, method, process, or assessment is established
for a dened purpose.

22
Bibliography
Alépée N., Bahinski T., Daneshian M., De Wever B., Fritsche E., Goldberg A., Hansmann J., Hartung T., Haycock J., Hogberg H.,
Hoelting L., Kelm J.M., Kadereit S., McVey E., Landsiedel R., Leist M., Lübberstedt M., Noor F., Pellevoisin C., Petersohn D.,
Pfannenbecker U., Reisinger K., Ramirez T., Rothen-Rutishauser B., Schäfer-Korting M., Zeilinger K. and Zurich M-G. (2014) “State-of-
the-art of 3D cultures (organs-on-a-chip) in safety testing and pathophysiology—a t4 report” ALTEX, 31:441-477.
Andersen M., Betts K., Dragan Y., Fitzpatrick S., Goodman J.L., Hartung T., Himmelfarb J., Ingber D.E., Jacobs A., Kavlock R., Kolaja K.,
Stevens J.L., Tagle D., Taylor D.L. and Throckmorton D. (2014) “Developing microphysiological systems for use as regulatory tools -
challenges and opportunities” ALTEX, 31:364-367.
Aung M.T., Yu Y., Ferguson K.K., Cantonwine D.E., Zeng L., McElrath T.F., Pennathur S., Mukherjee B., Meeker J.D. (2021) “Cross-
Sectional Estimation of Endogenous Biomarker Associations with Prenatal Phenols, Phthalates, Metals, and Polycyclic Aromatic
Hydrocarbons in Single-Pollutant and Mixtures Analysis Approaches” Environ Health Perspect, 129(3):37007. Epub 2021 Mar 24.
PMID: 33761273; PMCID: PMC7990518. doi: 10.1289/EHP7396
Bhatia, S.N. and Ingber, D.E. (2014) “Microuidic organs-on-chips. Nature biotechnology” 32, 760–772.
Blessinger T, Davis A, Chiu WA, Stanek J, Woodall GM, Gift J, Thayer KA, Bussard D. (2020) “Application of a unied probabilistic
framework to the dose-response assessment of acrolein.” Environ Int. 2020 Aug 5;143:105953. PMID: 32768806. doi: 10.1016/j.
envint.2020.105953.
Bopp S.K., Kienzler A., Richarz A., van der Linden S.C., Paini A., Parissis N. and Worth A.P. (2019) “Regulatory assessment and
risk management of chemical mixtures: challenges and ways forward” Critical Reviews in Toxicology, 49(2):174-189. DOI:
10.1080/10408444.2019.1579169
Bornmann, L., Haunschild, R. & Mutz, R. (2021) Growth rates of modern science: a latent piecewise growth curve approach to model
publication numbers from established and new literature databases. Humanit Soc Sci Commun 8, 224. https://doi.org/10.1057/
s41599-021-00903-w
Burnett S., Blanchette A., Chiu S.A. and Rusyn, I. (2021) “Human induced pluripotent stem cell (iPSC)-derived cardiomyocytes as an in
vitro model in toxicology: strengths and weaknesses for hazard identication and risk characterization” Expert Opinion on Drug
Metabolism and Toxicology, 17:8, 887-902, DOI: 10.1080/17425255.2021.1894122
Chiu W.A. (2017) “Chemical risk assessment and translation to socio-economic assessments” OECD Environment Working Papers,
March 14; (117). http://dx.doi.org/10.1787/a930054b-en
Chiu W.A., Axelrad D., Dalaijamts C., Dockins C., Shao K., Shapiro A.J., and Paoli G. (2018) “Beyond the RfD: Broad Application
of a Probabilistic Approach to Improve Chemical Dose-Response Assessments for Noncancer Effects.” EHP, DOI: https://doi.
org/10.1289/ehp3368
Chiu, W. A., Ginsberg, G.L., and Pullen, K. (2010) “The Exposome: A Powerful Approach for Evaluating Environmental Exposures and
Their Inuences on Human Disease”
Chiu W.A. and Paoli G.M. (2020) “Recent Advances in Probabilistic Dose-Response Assessment to Inform Risk-Based Decision Making.”
Risk Anal. PMID: 32966629 doi: 10.1111/risa.13595.
Chiu W.A. and Rusyn I. (2018) “Advancing chemical risk assessment decision-making with population variability data: challenges and
opportunities” Mamm Genome, doi: 10.1007/s00335-017-9731-6. [Epub ahead of print] PubMed PMID: 29299621
Chiu W.A. and Slob W. (2015): “A Unied Probabilistic Framework for Dose-Response Assessment of Human Health Effects” EHP, DOI:
http://dx.doi.org/10.1289/ehp.1409385
Clarke G.A., Hartse B.X., Asli A.E.N., Taghavimehr M., Hashemi N., Shirsavar M.A., Montazami R., Alimoradi N., Nasirian V., Ouedraogo
L.J., and Hashemi N.N. (2021) “Advancement of Sensor Integrated Organ-on-Chip Devices” Sensors 21, 1367

23
Densen P. (2011) "Challenges and opportunities facing medical education." Trans Am Clin Climatol Assoc. 122:48-58. PMID: 21686208;
PMCID: PMC3116346.
Dennis K.K., Marder E., Balshaw D.M., Cui Y., Lynes M., Patti G.J., Rappaport S.M., Shaughnessy D.T., Vrijheid M., and Barr D.B. (2017)
“Biomonitoring in the era of the exposome” Environ Health Perspect 125, 502-510. doi:10.1289/EHP474
Environmental Protection Agency. (2021) “A Review of U.S. EPA’s ORD Staff Handbook for Developing IRIS Assessments: 2020 Version”
Washington, DC: The National Academies Press, https://doi.org/10.17226/26289.
European Food Safety Authority and Evidence-Based Toxicology Collaboration. (2018) “Evidence integration in risk assessment:
The science of combining apples and oranges” EFSA Scientic Colloquium 23, Supporting Publication 16, EN-1396. doi:10.2903/
sp.efsa.2018.EN-1396
Escher B.I., Stapleton H.M. and Schymanski E.L. (2020) “Tracking complex mixtures of chemicals in our changing environment” Science,
80 (367), 388-392. 10.1126/science.aay6636
Fantke P., Aylward L., Bare J., Chiu W.A., Dodson R., Dwyer R., Ernstoff A., Howard B., Jantunen M., Jolliet O., Judson R., Kirchhübel N.,
Li D., Miller A., Paoli G., Price P., Rhomberg L., Shen B., Shin H-M., Teeguarden J., Vallero D., Wambaugh J., Wetmore B.A., Zaleski
R., and McKone T.E. (2018) “Advancements in Life Cycle Human Exposure and Toxicity Characterization” Environ Health Perspect,
126(12):125001. doi: 10.1289/EHP3871. PubMed PMID: 30540492.
Fantke P., Chiu W.A., Aylward L., Judson R., Huang L., Jang S., Gouin T., Rhomberg L., Aurisano N., McKone T. and Jolliet O. (2021)
“Exposure and Toxicity Characterization of Chemical Emissions and Chemicals in Products: Global Recommendations and
Implementation in USEtox” Int J Life Cycle Assess, 26(5):899-915. doi: 10.1007/s11367-021-01889-y. Epub 2021 Apr 5. PMID:
34140756; PMCID: PMC8208704.
Farhat N., Tsaioun K., Saunders-Hastings P., Morgan R.L., Ramoju S., Hartung T., Krewski D. (2022) “Systematic review in evidence-based
risk assessment” ALTEX, 39(3), 463–479. doi: 10.14573/altex.2004111
Ferrario D., Brustio R. and Hartung T. (2014) “Glossary of reference terms for alternative test methods and their validation” ALTEX,
31:319-335.
Fetah K., Tebon P., Goudie M.J., Eichenbaum J., Ren L., Barros N., Nasiri R., Ahadian S., Ashammakhi N., Dokmeci M.R., and
Khademhosseini A. (2019) “The emergence of 3D bioprinting in organ-on-chip systems” Prog Biomed Eng 1, 012001.
Ginsberg G.L., Fedinick K.P., Solomon G.M., Elliott K.C., Vandenberg J.J., Barone S., and Bucher J.R. (2019) “New Toxicology Tools and
the Emerging Paradigm Shift in Environmental Health Decision-Making” Environ Health Perspect, 127(12):125002. doi: 10.1289/
EHP4745. Epub 2019 Dec 13. PMID: 31834829; PMCID: PMC6957281.
Hartung T. (2009a) “Toxicology for the twenty-rst century” Nature, 460:208-212. doi:10.1038/460208a
Hartung T. (2009b) “Toxicology for the 21st century: Mapping the road ahead” Tox. Sci., 109:18-23.
Hartung T. (2017a) “Evolution of toxicological science: the need for change” International Journal of Risk Assessment and
Management, 20:21-45.
Hartung T. (2017b) “Thresholds of Toxicological Concern—setting a threshold for testing where there is little concern” ALTEX, 34:331-
351. doi: 10.14573/altex.1707011
Hartung, T. (2017c) “Utility of the adverse outcome pathway concept in drug development” Expert Opin Drug Metab Toxicol 13, 1-3. do
i:10.1080/17425255.2017.1246535
Hartung T. (2018) “Perspectives on in vitro to in vivo extrapolations” Journal of Applied In vitro Toxicology, 4:305–316. Doi: 10.1089/
aivt.2016.0026
Hartung T. and McBride M. (2011) “Food for thought… on mapping the human toxome” ALTEX, 28, 83-93. doi: 10.14573/
altex.2011.2.083

24
Hartung, T. and Leist, M. (2008) “Food for thought … on the evolution of toxicology and phasing out of animal testing” ALTEX 25, 91-
96. doi:10.14573/altex.2008.2.91
Hartung T. and Rovida C. (2009) “Chemical regulators have overreached” Nature, 460:1080-1081.
Hernández A.F. and Tsatsakis A.M. (2017) “Human exposure to chemical mixtures: Challenges for the integration of toxicology with
epidemiology data in risk assessment” Food and Chemical Toxicology, 103: 188-193. https://doi.org/10.1016/j.fct.2017.03.012
Hoffmann S. and Hartung T. (2005) “Diagnosis: Toxic!—Trying to apply approaches of clinical diagnostics and prevalence in toxicology
considerations” Toxicol. Sci, 85, 422-428. doi: 10.1093/toxsci/k099
Hoffmann S., de Vries R.B.M., Stephens M.L., Beck N.B., Dirven H.A.A.M., Fowle J.R., Goodman J.E., Hartung T., Kimber I., Lalu M.M.,
Thayer K., Whaley P., Wikoff D., and Tsaioun K. (2017) “A primer on systematic reviews in toxicology” Arch Toxicol 91, 2551-2575.
doi:10.1007/s00204-017-1980-3
Hood, L. and Rowen, L. (2013) “The Human Genome Project: big science transforms biology and medicine” Genome Med 5, 79.
https://doi.org/10.1186/gm483
Huang H., Wang A., Morello-Frosch R., Lam J., Sirota M., Padula A., and Woodruff T.J. (2018) “Cumulative Risk and Impact Modeling on
Environmental Chemical and Social Stressors” Curr Envir Health Rpt 5, 88–99. https://doi.org/10.1007/s40572-018-0180-5
Kleensang A., Maertens A., Rosenberg M., Fitzpatrick S., Lamb J., Auerbach S., Brennan R., Crofton K.M., Gordon B., Fornace A.J.
Jr., Gaido K., Gerhold D., Haw R., Henney A., Ma’ayan A., McBride M., Monti S., Ochs M.F., Pandey A., Sharan R., Stierum R.,
Tugendreich S., Willett C., Wittwehr C., Xia J., Patton G.W., Arvidson K., Bouhifd M., Hogberg H.T., Luechtefeld T., Smirnova L., Zhao
L., Adeleye Y., Kanehisa M., Carmichael P., Andersen E.M., Hartung T. (2014) “Pathways of Toxicity” ALTEX, 31:53-61. doi: 10.14573/
altex.1309261
Kvasnicka J., Stylianou K.S., Nguyen V.K., Huang L., Chiu W.A., Burton G.A., Semrau J., and Jolliet O. (2019) “Human Health Benets
from Fish Consumption vs. Risks from Inhalation Exposures Associated with Contaminated Sediment Remediation: Dredging of
the Hudson River” Environ Health Perspect, 127(12):127004. doi:10.1289/EHP5034. Epub 2019 Dec 13. PubMed PMID: 31834828;
PubMed Central PMCID: PMC6957280.
Krewski D., Andersen M., Tyshenko M.G., Krishnan K., Hartung T., Boekelheide K., Wambaugh J.F., Jones D., Whelan M., Thomas
R., Yauk C., Barton-Maclaren T. and Cote I. (2020) “Toxicity Testing in the 21st Century: Progress in the past decade and future
perspectives” Arch Toxicol, 94:1–58.
Krewski, D., Saunders-Hastings, P., and Baan, R., et al. (2022) “Workshop Report: Development of an evidence-based risk assessment
framework” ALTEX, in press.
Krewski, D., Saunders-Hastings, P., and Arzuga, X., et al. (in preparation) “Development of a framework for evidence synthesis:
Workshop report”
Lakhani, C. M., et al. (2019) “Repurposing large health insurance claims data to estimate genetic and environmental contributions in 560
phenotypes” Nat Genet 51, 327–334.
Lynn R. Goldman. (2003) “The Red Book: A Reassessment of Risk Assessment, Human and Ecological Risk Assessment: An International
Journal” 9:5, 1273-1281. https://doi.org/10.1080/10807030390248492
Materne E-M., Tonevitsky A.G., and Marx U. (2013) “Chip-based liver equivalents for toxicity testing—organotypicalness versus cost-
efcient high throughput” Lab Chip, 13, 3481 DOI: 10.1039/C3LC50240F
Maertens A., Golden E., Luechtefeld T.H., Hoffmann S., Tsaioun K. and Hartung T. (2022) “Probabilistic Risk Assessment—the Keystone
for the Future of Toxicology” ALTEX, 39:3-29. doi:10.1,4573/altex.2201081
Marx U., Akabane T., Andersson T.B., Baker E., Beilmann M., Beken S., Brendler-Schwaab S., Cirit M., David R., Dehne E-M., Durieux I.,
Ewart L., Fitzpatrick S.C., Frey O., Fuchs F., Grifth L.G., Hamilton G.A., Hartung T., Hoeng J., Hogberg H., Hughes D.J., Ingber D.E.,
Iskandar A., Kanamori T., Kojima H., Kuehnl J., Leist M., Li B., Loskill P., Mendrick D.L., Neumann T., Pallocca G., Rusyn I., Smirnova

25
L., Steger-Hartmann T., Tagle D.A., Tonevitsky A., Tsyb S., Trapecar M., van de Water B., van den Eijnden-van Raaij J., Vulto P.,
Watanabe K., Wolf A., Zhou X. and Roth A. (2020) “Biology-inspired microphysiological systems to advance medicines for patient
benet and animal welfare” ALTEX, 37:364-394. doi: 10.14573/altex.2001241
Marx U., Andersson T.B., Bahinski A., Beilmann M., Beken S., Cassee F.R., Cirit M., Daneshian M., Fitzpatrick S., Frey O., Gaertner C.,
Giese C., Grifth L., Hartung T., Heringa M.B., Hoeng J., de Jong W.H., Kojima H., Kuehnl J., Luch A., Maschmeyer I., Sakharov D.,
Sips A.J.A.M., Steger-Hartmann T., Tagle D.A., Tonevitsky A., Tralau T., Tsyb S., van de Stolpe A., Vandebriel R., Vulto P., Wang J.,
Wiest J., Rodenburg M. and Roth A. (2016) “Biology-inspired microphysiological system approaches to solve the prediction dilemma
of substance testing using animals” ALTEX, 33:272-321. doi: 10.14573/altex.1603161
Meigs L., Smirnova L., Rovida C., Leist M., and Hartung T. (2018) “Animal testing and its alternatives—the most important omics is
economics” ALTEX, 35:275-305. doi: 10.14573/altex.1807041
National Academies of Sciences, Engineering, and Medicine. (2009) “Science and Decisions: Advancing Risk Assessment” Washington,
DC: The National Academies Press. https://doi.org/10.17226/12209
National Academies of Sciences, Engineering, and Medicine. (2011) “Review of the Environmental Protection Agency’s Draft IRIS
Assessment of Formaldehyde” Washington, DC: The National Academies Press. https://doi.org/10.17226/13142
National Academies of Sciences, Engineering, and Medicine. (2016) “Interindividual Variability: New Ways to Study and Implications for
Decision Making: Workshop in Brief” Washington, DC: The National Academies Press. https://doi.org/10.17226/23413
National Academies of Sciences, Engineering, and Medicine. (2017b) “Application of Systematic Review Methods in an Overall Strategy
for Evaluating Low-Dose Toxicity from Endocrine Active Chemicals” Washington, DC: The National Academies Press, https://doi.
org/10.17226/24758.
National Academies of Sciences, Engineering, and Medicine. (2017a) “Using 21st Century Science to Improve Risk-Related Evaluations”
Washington, DC: The National Academies Press. https://doi.org/10.17226/24635
National Academies of Sciences, Engineering, and Medicine. (2021) “Microphysiological Systems: Bridging Human and Animal
Research: Proceedings of a Workshop—in Brief” Washington, DC: The National Academies Press. https://doi.org/10.17226/26124
National Research Council. (1983) “Risk Assessment in the Federal Government: Managing the Process” Washington, DC: The National
Academies Press. https://doi.org/10.17226/366
National Research Council. (2007) “Toxicity Testing in the 21st Century: A Vision and a Strategy” Washington, DC: The National
Academies Press. https://doi.org/10.17226/11970
National Research Council. (2012) “Exposure Science in the 21st Century: A Vision and a Strategy” Washington, DC: The National
Academies Press. https://doi.org/10.17226/13507
Nitsche K.S., Müller I., Malcomber S., Carmichael P.L. and Bouwmeester H. (2022) “Implementing organ-on-chip in a next-generation
risk assessment of chemicals: a review” Arch Toxicol 96, 711–741.
Pamies D., Leist M., Coecke S., Bowe G., Allen D., Gstraunthaler G., Bal-Price A., Pistollato F., DeVries R., Hogberg H.T., Hartung T. and
Stacey G. (2022) “Guidance Document on Good Cell and Tissue Culture Practice 2.0 (GCCP 2.0)” ALTEX, 39:30–70. doi: 10.14573/
altex.2111011
Partosch F., Mielke H., Stahlmann R., Kleuser B., Barlow S., and Gundert-Remy U. (2015). “Internal threshold of toxicological concern
values: Enabling route-to-route extrapolation: Arch Toxicol 89, 941-948. doi:10.1007/s00204- 014-1287-6
Poutasse C.M., Herbstman J.B., Peterson M.E., Gordon J., Soboroff P.H., Holmes D., Gonzalez D., Tidwell L.G., and Anderson K.A.
(2019) “Silicone Pet Tags Associate Tris (1,3-dichloro-2-isopropyl) Phosphate Exposures with Feline Hyperthyroidism” Environ Sci
Technol. 53(15):9203-9213. doi: 10.1021/acs.est.9b02226. Epub 2019 Jul 10. PMID: 31290326; PMCID: PMC7330886
Rappaport S.M. and Smith M.T. (2010) “Environment and Disease Risks” Science, 330: 460–461. doi:10.1126/science.1192603
Richard A.M., Huang R., Waidyanatha S., Shinn P., Collins B.J., Thillainadarajah I., Grulke C.M., Williams A.J., Lougee R.R., Judson
26
R.S., Houck K.A., Shobair M., Yang C., Rathman J.F., Yasgar A., Fitzpatrick S.C., Simeonov A., Thomas R.S., Crofton K.M., Paules
R.S., Bucher J.R., Austin C.P., Kavlock R.J., Tice R.R. (2021) “The Tox21 10K Compound Library: Collaborative Chemistry Advancing
Toxicology” Chem Res Toxicol. 34(2):189-216. doi: 10.1021/acs.chemrestox.0c00264. Epub 2020 Nov 3. PMID: 33140634; PMCID:
PMC7887805
Robinson, O., and Vrijheid, M. (2015) “The Pregnancy Exposome” Curr Envir Health Rpt 2, 204–213. https://doi.org/10.1007/s40572-015-
0043-2
Roth A., M.P.S.-W.S. Berlin, Marx U., Vilén L., Ewart L., Grifth L.G., Hartung T., Ingber D.E., Mendrick D.L., Steger-Hartmann T., and
Tagle D.A. (2021) “2019 Human microphysiological systems for drug development” Science, 373:1304-1306
Rusyn I., Chiu W.A., and Wright F.A. (2022) “Model systems and organisms for addressing inter- and intra-species variability in risk
assessment.” Regul Toxicol Pharmacol. 2022 May 27:105197. doi: 10.1016/j.yrtph.2022.105197. Epub ahead of print. PMID: 35636685.
Samet J.M., et al. (2020) “The IARC Monographs: Updated procedures for modern and transparent evidence synthesis in cancer hazard
identication” Journal of the National Cancer Institute, 112(1):30-37. 10.1093/jnci/djz169
Sillé F.C.M., Karakitsios S., Kleensang A., Koehler K., Maertens A., Miller G.W., Prasse C., Quiros-Alcala L., Ramachandran G., Rappaport
S.M., Rule A.M., Sarigiannis D., Smirnova L., and Hartung T. (2020) “The exposome—a new approach for risk assessment” ALTEX, 37:
3-23. doi: 10.14573/altex.2001051
van Vliet E., Daneshian M., Beilmann M., Davies A., Fava E., Fleck R., Julé Y., Kansy M., Kustermann S., Macko P., Mundy W., Roth A.,
Shah I., Uteng M., van de Water B., Hartung T. and Leist M. (2014) “Current approaches and future role of high content imaging in
safety sciences and drug discovery” ALTEX, 31:479-493.
Vermeulen R., Schymanski E.L., Barabási A.L., and Miller, G.W. (2020) “The exposome and health: Where chemistry meets biology”
Science, 367(6476):392-396. doi: 10.1126/science.aay3164. PMID: 31974245; PMCID: PMC7227413
Wambaugh J.F., Wetmore B.A., Pearce R., Strope C., Goldsmith R., Sluka J.P., Sedykh A., Tropsha A., Bosgra S., Shah I., Judson R.,
Thomas R.S., and Setzer R.W. (2015) “Toxicokinetic triage for environmental chemicals” Toxicol Sci 147, 55-67. doi:10.1093/toxsci/
kfv118
Wang, Z., Walker, G. W., Muir, D. C. G., and Nagatani-Yoshida, K. (2020) “Toward a global understanding of chemical pollution: a rst
comprehensive analysis of national and regional chemical inventories,” Environ Sci Technol 54, 2575–2584. http://doi.org/10.1021/
acs.est.9b06379
World Health Organization & International Programme on Chemical Safety. (2018) “Guidance document on evaluating and expressing
uncertainty in hazard characterization” 2nd ed. World Health Organization. https://apps.who.int/iris/handle/10665/259858. License:
CC BY-NC-SA 3.0 IGO
Wild, C.P. (2019) “The global cancer burden: necessity is the mother of prevention” Nature Reviews Cancer 19: 123-124. https://doi.
org/10.1038/s41568-019-0110-3
Wilkinson M.D., Dumontier M., Aalbersberg I.J., Appleton G., Axton M., Baak A., Blomberg N., Boiten J-W., da Silva Santos L.B.,
Bourne P.E., Bouwman J., Brookes A.J., Clark T., Crosas M., Dillo I., Dumon O., Edmunds S., Evelo C.T., Finkers R., Gonzalez-Beltran
A., Gray A.J.G., Growth P., Goble C., Grethe J.S., Heringa J., Hoen P.A.C., Hooft R., Kuhn T., Kok R, Kok J., Lusher S.J., Martone M.E.,
Mons A., Packer A.L., Persson B., Rocca-Serra P., Roos M., van Schaik R., Sansone A-A., Schultes E., Sengstag T., Slater T., Strawn
G., Swertz M.A., Thompson M., van der Lei J., van Mulligen E., Velterop J., Waagmeester A., Wittenburg P., Wolstencroft K., Zhao J.
and Mons B. (2016) “The FAIR Guiding Principles for scientic data management and stewardship” Sci Data 3, 160018. https://doi.
org/10.1038/sdata.2016.18
Woodruff T.J., Sutton P. (2014) “The Navigation Guide systematic review methodology: A rigorous and transparent method for
translating environmental health science into better health outcomes” Environmental Health Perspectives, 122(10):1007-1014.
27
Appendix I—Workshop Attendees
Workshop Co-chairs
Ana Navas-Acien Columbia University
Weihsueh Chiu Texas A&M University
Thomas Hartung Johns Hopkins University
Workshop Participants
Tony Atala WakeForest University
Dana Dolinoy University of Michigan
Lauren Heine ChemForward
Salman Khetani University of Illinois at Chicago
Marianthi Kioumortzoglu Columbia University
Nicole Kleinstreuer NIEHS NTP
Koren Mann McGill University
Uwe Marx TissUse
Patrick McMullen Scitovation
Gary Miller Columbia University
Katie Paul- Friedman US Environmental Protection Agency
Jennifer Sass NRDC
Kris Thayer US Environmental Protection Agency
Cavin Ward-Caviness US Environmental Protection Agency
Cheryl Walker Baylor University
Katrina Waters Pacic Northwest National Laboratory
Hao Zhu Rutgers University
Government Observers
Bindu Nair OUSD(R&E), Basic Research Ofce
Jean Luc Cambrier OUSD(R&E), Basic Research Ofce
Shanni Silberberg OUSD(R&E), Basic Research Ofce
Daniel Osburn OUSD(R&E), Basic Research Ofce
Betsy Melebrink OUSD(R&E), Basic Research Ofce
Anna Lowit Environmental Protection Agency
Mark Johnson US Army Public Health Center
Rabih Jabbour US Army Edgewood Chemical Biological Center
Natalie Vinas US Army Engineer Research and Development Center
Louis Scarano Environmental Protection Agency
Rachel Gooding Department of Homeland Security, Chemical Security Analysis Center
VT-ARC Team
Matthew Bigman Virginia Tech Applied Research Corporation
Jordan Brown Virginia Tech Applied Research Corporation
Christina Houfek Virginia Tech Applied Research Corporation
Kate Klemic Virginia Tech Applied Research Corporation
Lynne Ostrer Virginia Tech Applied Research Corporation

28
Workshop Participant Short Biography
Anthony Atala, Director, Wake Forest Institute for Regenerative Medicine
Wake Forest School of Medicine
Anthony Atala, MD, is the G. Link Professor and Director of the Wake Forest Institute for Regenerative
Medicine, and the W. Boyce Professor and Chair of the Department of Urology at Wake Forest University.
His work focuses on growing human cells, tissues and organs. Fifteen applications of technologies
developed in Dr. Atala’s laboratory have been used clinically. Dr. Atala was named by Scientic American
as one of the world’s most inuential people in biotechnology, by U.S. News and World Report as one
of 14 Pioneers of Medical Progress in the 21st Century, by Life Sciences Intellectual Property Review as
one of 50 key inuencers in the life sciences intellectual property arena, and by the journal Nature Biotechnology as one of the top 10
translational researchers in the world.
Weihsueh A. Chiu, Professor
Texas A&M University
Weihsueh A. Chiu, Ph.D. is a professor in the Department of Veterinary Physiology and Pharmacology
at Texas A&M University. Before joining the university in 2015, he worked at the U.S. Environmental
Protection Agency (EPA) for more than 14 years, most recently as branch chief in the Ofce or Research
and Development. His research in human health risk assessment includes toxicokinetics, physiologically-
based pharmacokinetic modeling, dose-response assessment, characterizing uncertainty and variability,
systematic review, and meta-analysis, with particular interest in Bayesian and probabilistic methods.
Dr. Chiu has participated or chaired expert review panels for multiple government agencies, including NTP, CalEPA, the FDA, and
ATSDR. He has also served on numerous national and international committees and workgroups for Health Canada, the World
Health Organization, the Organisation for Economic Cooperation and Development, and the U.S. National Academies of Sciences,
Engineering and Medicine.
Dana Dolinoy, Chair and Professor
University of Michigan
Dana C. Dolinoy is Professor of Environmental Health Sciences and Nutritional Sciences and NSF
International Chair of Environmental Health Sciences at the University of Michigan School of Public Health
as well as Faculty Director of the Epigenomics Core at Michigan Medicine. Her research focuses on
how nutritional and environmental factors interact with epigenetic gene regulation to shape health and
disease. In 2015, she received the 2015 NIH Director’s Transformative Research Award to develop piRNA
epigenetic editing technologies and in 2018 received the Society of Toxicology Achievement Award and
has recently co-edited the book ToxicoEpigenetics: Core Principles and Applications. She has authored
>130 manuscripts and 10 book chapters, and served as Chair of the Gordon Conference in Cellular and
Molecular Mechanisms of Toxicity. She has mentored 14 doctoral students, one of whom recently received
a F31 award, and 6 post-doctoral fellows, one of whom recently received a NIEHS K99/R00 award, as well as several masters and
undergraduate students.
Thomas Hartung, Professor and Chair
Johns Hopkins University
https://www.jhsph.edu/faculty/directory/prole/2308/thomas-hartung
Thomas Hartung is the Doerenkamp-Zbinden Chair for Evidence-based Toxicology in the Department of
Environmental Health and Engineering at Johns Hopkins Bloomberg School of Public Health, Baltimore,
with a joint appointment at the Whiting School of Engineering. He also holds a joint appointment for
Molecular Microbiology and Immunology at the Bloomberg School. He is adjunct afliate professor
at Georgetown University, Washington D.C.. In addition, he holds a joint appointment as Professor for
Pharmacology and Toxicology at the University of Konstanz, Germany; he also is Director of Centers for Alternatives to Animal Testing
(CAAT, http://caat.jhsph.edu) of both universities.

29
CAAT hosts the secretariat of the Evidence-based Toxicology Collaboration (http://www.ebtox.org), the Good Read-Across Practice
Collaboration, the Good Cell Culture Practice Collaboration, the Green Toxicology Collaboration and the Industry Renement Working
Group. As PI, Dr. Hartung headed the Human Toxome project funded as an NIH Transformative Research Grant and the series of World
Summits for Microphysiological Systems started in 2022. He is Field Chief Editor of Frontiers in Articial Intelligence. He is the former
Head of the European Commission’s Center for the Validation of Alternative Methods (ECVAM), Ispra, Italy, and has authored more than
625 scientic publications (h-index 105).
Lauren Heine, Director of Science and Data Integrity
ChemFORWARD
www.chemforward.org
lauren@chemforward.org
Lauren Heine applies green chemistry, green engineering, alternatives assessment and multi-stakeholder
collaboration to develop tools that result in safer and more sustainable chemical products and processes.
Her work with ChemFORWARD builds on prior experience developing GreenScreen’s for Safer
Chemicals, a pioneering method for chemical hazard assessment to enable informed substitution; and
CleanGredients, a web-based information platform for identifying greener chemicals for use in cleaning
products; both tools were designed to scale access to information needed to develop materials and products that are safe and circular.
Lauren worked closely with the US EPA Safer Choice Program to facilitate development of ingredient and hazard criteria for the Safer
Choice Program.
Salman Khetani, Associate Professor and Director of Graduate Studies
University of Illinois at Chicago
mtm.uic.edu
Salman Khetani is an associate professor of Biomedical Engineering at the University of Illinois at Chicago
where he directs the Microfabricated Tissue Models (MTM) laboratory that is engaged in developing in
vitro models of various tissues (liver, cardiac, intestine, brain, and placenta) for drug screening, disease
modeling, and regenerative medicine. Prior to academia, Dr. Khetani co-founded and directed research
at Hepregen Corporation, which launched engineered models of the human liver that continue to serve
the pharmaceutical industry for elucidating drug metabolism, toxicity, and efcacy for liver diseases. Dr.
Khetani’s research is currently funded by the US National Science Foundation and National Institutes of Health. His laboratory focuses
on engineering the microenvironmental cues around mammalian cells towards stabilizing their long-term phenotype in vitro for
applications in drug development and regenerative medicine. He has developed model systems to mimic key aspects of diseases in
vitro and elucidate underlying molecular mechanisms of disease progression as a function of cell-cell, cell-ECM, and cell-soluble factor
interactions. His liver models have been translated to the commercial realm through licensing of issued patents and patent applications
to companies. Recent work on iPSC-derived atrial cardiomyocytes are used to study the underlying genetic determinants of atrial
brillation in close collaboration with leading cardiologists.
Marianthi-Anna Kioumourtzoglou, Assistant Professor
Columbia University
https://www.publichealth.columbia.edu/people/our-faculty/mk3961
Marianthi-Anna Kioumourtzoglou is an environmental engineer and epidemiologist. She holds a Master
of Science in Public Health (MSPH) from the Environmental Sciences and Engineering Department at the
University of North Carolina at Chapel Hill and a Doctor of Science (ScD) in Environmental Health from
the Harvard TH Chan School of Public Health, where she also conducted her post-doctoral fellowship.
She is currently an Assistant Professor at the Department of Environmental Health Sciences at Columbia
University’s Mailman School of Public Health. Her research focuses on applied statistical issues related
to environmental epidemiology, including quantifying and correcting for exposure measurement
error, exposure prediction uncertainty propagation, and assessment of high-dimensional and complex
exposures in health analyses. Her studies mainly (albeit not exclusively) focus on air pollution exposures
and, additionally, on identifying vulnerable sub-populations and characterizing how risks may vary across neighborhood-level and other
urban characteristics, as well as in a changing climate.

30
Nicole Kleinstreuer, Director
NICEATM
https://www.publichealth.columbia.edu/people/our-faculty/mk3961
Nicole Kleinstreuer is the acting director of the NTP Interagency Center for the Evaluation of Alternative
Toxicological Methods (NICEATM), the US federal resource for alternatives to animal testing. At NICEATM,
she leads domestic and international efforts to develop novel testing and analysis strategies that provide
more rapid, mechanistic, and human-relevant predictions of potential environmental chemical hazards.
Kleinstreuer’s research focuses on mathematical and computational modeling of biological systems and
their susceptibility to perturbations that result in adverse health outcomes. She has a secondary appointment in the NIEHS Division
of Intramural Research Biostatistics and Computational Biology Branch, and adjunct faculty positions in the Yale University School of
Public Health and the Eshelman School of Pharmacy at UNC Chapel Hill.
Koren Mann, Professor and Chair
McGill University, Department of Pharmacology and Therapeutics
Koren Mann is a Professor and Chair of the Department of Pharmacology and Therapeutics at McGill
University, and a Senior Investigator at the Lady Davis Institute for Medical Research in Montreal, Quebec,
Canada. She received her doctorate in Pathology and Immunology from Boston University in 1999 and was
a postdoctoral fellow at McGill University from 1999-2004. Her laboratory focuses on studying the health
effects of metal exposure, although recent studies include other environmental pollutants. She focuses on
how modulation of the immune system results in pathology, especially cardiovascular toxicity. The overarching theme of her lab is to
integrate toxicology questions within the framework of the epidemiology, providing relevance and feed-forward questions to further
interrogate in human cohorts.
Uwe Marx, MD
TissUse GmbH
www.tissuse.com
Uwe Marx is the founder and CSO of TissUse, a 2010 spin-out from the Technische Universitat Berlin
dedicated to the development of human organ and body-on-a-chip systems for drug testing and precision
medicine approaches. The solutions aim to shorten the drug development process and to reduce animal
experiments. With more than 30 years of experience in protein drug development and tissue engineering
Dr. Marx has published about 150 scientic papers and numerous reviews and book chapters. He is an inventor in more than 30 patent
families. Uwe Marx received his doctorate degree in immunology from the Charite of the Humboldt-University in Berlin in 1991 after
nishing his medical and biochemistry training. His academic research at the Charite Berlin, the University of Leipzig and the Technische
Universitat Berlin focused on human monoclonal antibodies, tissue engineering and human multi-organ chip solutions respectively.
Between 2000 and 2010, Uwe Marx joined ProBioGen, a biotech Company he founded in 1994 - as CSO. He served as a reviewer for
various German governmental biotech programmes and received several awards for the development of animal-free technologies. Dr.
Marx is a serial entrepreneur and co-founder of numerous German biotech companies.
Patrick McMullen, Director of Computational Toxicology
ScitoVation
www.scitovation.com
Patrick McMullen is the Director of Computational Toxicology at ScitoVation. Dr. McMullen works with
diverse stakeholders spanning government, non-prot, and industry groups to bring new approaches
to toxicology, with the goal of improving chemical safety decision-making processes. His research
and consulting work combine high-content biological experiments with statistical and computational
approaches to advance the understanding of biological fundamentals that underlie chemical safety
challenges. Dr. McMullen’s background in molecular biology, engineering, and computational science has been instrumental in
interpreting and communicating complex data problems in diverse applications. Dr. McMullen manages a diverse computational
biology team that uses modeling and cell-based experiments to deepen our understanding of how chemicals interact with biological
systems. Dr. McMullen earned his Ph.D. in Chemical and Biological Engineering from Northwestern.

31
Gary Miller, Vice Dean for Research Strategy and Innovation
Columbia University
https://www.publichealth.columbia.edu/people/our-faculty/gm2815
Gary Miller serves as Vice Dean for Research Strategy and Innovation and Professor of Environmental
Health Sciences at the Columbia University Mailman School of Public Health. He also has appointments
in the Department of Molecular Pharmacology and Therapeutics and the Department of Neurology in
the Vagelos College of Physicians and Surgeons. He is an international leader on the exposome, the
environmental analogue to the genome. Dr. Miller founded the rst exposome center in the U.S. and
wrote the rst book on the topic. He has helped develop high-resolution mass spectrometry methods
to provide an omic-scale analysis of the human exposome. He served as Editor-in-Chief of Toxicological
Sciences, the ofcial journal of the Society of Toxicology from 2013-2019 and is now Editor-in-Chief of
Exposome, the rst journal in the eld. He is on the Scientic Advisory Board of NIHs All of Us Research Program, the NIH Human
Health Exposure Analysis Resource (HHEAR), and the Human Biomonitoring for the European Union (HBM4EU) project.
Ana Navas-Acien, Professor
Columbia University
https://www.publichealth.columbia.edu/people/our-faculty/an2737
Ana Navas-Acien is a physician-epidemiologist (MD, University of Granada, Spain ‘96) with a specialty in
Preventive Medicine and Public Health (Hospital La Paz, Madrid ‘01) and a PhD in Epidemiology (Johns
Hopkins University ‘05). Her research investigates the long-term health effects of environmental exposures
(arsenic and other metals, tobacco smoke, e-cigarettes, air pollution), relevant molecular pathways, and
effective interventions for reducing involuntary exposures. She collaborates with major cohort studies such
as the Strong Heart Study, a study of cardiovascular disease and its risk factors in American Indian communities, and the Multi-Ethnic
Study of Atherosclerosis (MESA), a study of cardiovascular, metabolic and lung disease in urban settings across the US, and with the
TACT2 study (a clinical trial assessing the cardiovascular benets of metal chelation). Both in the US and internationally, she evaluates
exposure to tobacco smoke including e-cigarettes through toxicological and epidemiological research strategies. Her goals are to
contribute to the reduction of environmental health disparities in underserved and disproportionately exposed populations.
Katie Paul-Friedman, Toxicologist
US EPA
Dr. Katie Paul Friedman joined the Center for Computational Toxicology and Exposure in the Ofce of Research
and Development at the US EPA in August 2016, where she is currently focused on application of new approach
methodologies to chemical safety assessment, with additional interests in uncertainty in alternative and traditional
toxicity information, endocrine bioactivity and developmental neurotoxicity prediction, and in vitro kinetics. One
of her roles in the Center is to run the ToxCast program. Previously, Dr. Paul Friedman worked as a regulatory
toxicologist at Bayer CropScience with specialties in neuro-, developmental and endocrine toxicity, and predictive
toxicology. She has been actively involved in multi-stakeholder projects to develop adverse outcome pathways,
alternative testing approaches, and the regulatory acceptance of new approach methodologies. Her laboratory background includes
development of high-throughput screening assays, the combined use of myriad in vitro and in vivo approaches, including receptor-
reporter and biochemical assays, primary hepatocyte cultures, and targeted animal testing paradigms, to investigate the human
relevance of thyroid and metabolic adverse outcome pathways using probe chemicals. Dr. Paul Friedman received a Ph.D. in Toxicology
from the University of North Carolina at Chapel Hill.
Jennifer Sass, Senior Scientist
Natural Resources Defense Council
Jennifer Sass is a Senior Scientist at the Natural Resources Defense Council (2001-2021) and part-time
faculty at George Washington University Milken School of Public Health (2008-2021). She has published
over 50 articles. She holds BSc, MSc, and PhD (1998) degrees from the University of Saskatchewan,
College of Medicine, Department of Anatomy and Cell Biology, and a Post-Doctoral Certicate (2000)
from the University of Maryland, College of Medicine, Program in Human Health and the Environment.

32
Kristina Thayer, Director
US EPA, Chemical and Pollutant Assessment Division (CPAD)
thayer[email protected]
Kristina Thayer is Director of the Chemical and Pollutant Assessment Division (CPAD) at the U.S.
Environmental Protection Agency (https://www.epa.gov/aboutepa/about-chemical-and-pollutant-
assessment-division-cpad). CPAD occupies an essential position in EPA’s Ofce of Research and
Development between researchers generating scientic data and EPA’s program and regional ofces
that make decisions regarding the protection of public health and the environment. CPAD scientists
develop a range of t-for-purpose human health risk assessment products based on the evaluation,
synthesis, and analysis of the most up-to-date scientic information. Products include the Integrated Risk
Information System (IRIS) and Provisionally Peer Reviewed Toxicity Values (PPRTV) assessments. These
products are developed through interactions with EPA’s program and regional ofces, other agencies,
the scientic community, industry, policymakers, and the public. Once nalized, they serve as a major
scientic component supporting EPA’s regulations, advisories, policies, enforcement, and remedial action
decisions. CPAD also conducts cutting-edge research to develop innovative human health risk assessment methods (e.g., systematic
review) that facilitate careful evaluation of scientic evidence, as well as tools and models (e.g., benchmark dose modeling software).
Cheryl Walker, Director
Baylor College of Medicine, Center for Precision Environmental Health
Cheryl Walker holds the Alkek Presidential Chair in Environmental Health and is the founder and Director
of the Center for Precision Environmental Health at Baylor College of Medicine. She also directs the
NIEHS P30 Gulf Coast Center for Precision Environmental Health. Dr. Walker has over 200 publications in
the scientic literature and is an elected member of the National Academy of Medicine. Her research on
gene:environment interactions and environmental epigenomics has been continuously funded by the NIH,
DOD, and Foundations and advocacy groups for over 25 years Her laboratory actively investigates gene x
environment interactions and their role in diseases such as cancer, broids and NAFLD. The TSC2 tumor suppressor, and its role in cell
signaling, has been one of the areas of interest for her lab, in addition to the study of TSC2-linked pathways that regulate key cellular
functions.
Cavin Ward-Caviness, Computational Biologist
US EPA
Cavin Ward-Caviness is a Principal Investigator in the Public Health and Integrated Toxicology Division of the US
Environmental Protection Agency. With a background in computational biology and environmental epidemiology,
Dr. Ward-Caviness seeks to understand the environmental factors which inuence health in vulnerable populations
and the molecular mechanisms that inuence environmental health risks. He is the PI of the EPA CARES research
resource, which allows researchers to study environmental health effects in vulnerable patient populations, using
large electronic health record databases. Ward-Caviness also leads the Environmental Health Domain Team for the
National Covid Cohort Consortium. He is also interested in how epigenetics and metabolomics can serve as an
early indicator of adverse health effects from chemical and social environmental exposures and in particular how molecular biomarkers
can give us insight into how the environment may accelerate the aging process and thus contribute to chronic disease.
Katrina Waters, Director
Pacic Northwest National Laboratory
https://www.pnnl.gov/people/katrina-m-waters
Katrina Waters is a Laboratory Fellow and Director for Biological Sciences Research at the Pacic
Northwest National Laboratory (PNNL). Her research interests are focused at the intersection of
environmental exposures and infectious disease on human health. Her current programs include the study
of health effects of chemicals at Superfund sites and personal environmental exposure assessment for
epidemiological studies in disadvantaged communities. She recently completed a Department of Energy
research program focused on airborne and environmental transmission of COVID-19. She has also led
numerous research efforts in Computational Modeling, Bioinformatics, and Data Management for a NIAID
Center for Predictive Modeling of Infectious Diseases and a Department of Homeland Security program for Predictive Modeling of Viral
Infections. Dr. Waters holds joint faculty appointments with OSU and the University of Washington.

33
Hao Zhu, Professor
Rutgers University-Camden
Hao Zhu is a Professor of Chemistry at the Rutgers University-Camden. His major research interest is to
use cheminformatics tools to develop predictive models. All resulted models can be used to directly
predict the chemical toxicity based on the public big data and molecular structure information. His current
research interests also include data-driven modeling, articial intelligence algorithm development and
computer-aided nanomedicine design. He is the Principal Investigator of several prestigious research
grants (NIH R01, R15 and etc). Dr. Zhu is author/co-author of 81 peer-reviewed journal articles and 7 book chapters with over 5,600
citations. His research was recognized with different awards, such as Rutgers Chancellor’s Award for Outstanding Research and Creative
Activity, National Institute of Environmental Health Sciences (NIEHS) Extramural Paper of the Month (two times, 2019 and 2020) and
Drug Discovery Today top citation paper of the year.

34
Appendix II—Workshop Agenda and Prospectus
Basic Research Innovation Collaboration Center
4100 N. Fairfax Rd. | Fourth Floor| Suite 450
Arlington, VA 22203
DAY 1—THURSDAY, APRIL 28, 2022
Time Title Speaker
8:00—8:15 Check-in and Connental Breakfast
8:15 - 8:20 Welcome and Introducons and Expectaons Thomas Hartung, JHU
8:20 -8:45 Workshop Framing Talk Co-chairs
8:45—9:00
Breakout Instrucons and Morning Break
9:00—10:45
Working Group I: Dene the Problem
Small group discussions to frame a vision for toxicology as a predicve science and idenfy the
greatest hurdles to achieving it.
Group A—Exposure-driven Toxicology
Group B—Technical Advances
Group C—Evidence Integraon
10:45—11:00 BREAK - Transion to main conference room and leads prepare outbrieng
11:00 –12:00 Working Group 1: Outbrieng
12:00—1:00 LUNCH (provided for parcipants)
1:00—3:45
Working Group II: Technical Capabilies and Opportunies
What are the promising research direcons for moving to a more predicve
toxicology? What are the potenal capabilies in the 10- to 20-year horizon?
Group A—Exposure-driven Toxicology
Group B—Technical Advances
Group C—Evidence Integraon
3:45—4:00 BREAK - Transion to main room and leads prepare outbrieng
4:00—4:45 Working Group II: Outbrieng
4:45—5:00 Summary of Day Co-chairs
5:00 MEETING ADJOURNED FOR THE DAY
35
DAY 2—FRIDAY, APRIL 29
TH
, 2022
Time
Title Speaker
8:00—8:15 Check-in and Connental Breakfast
8:15—8:30 Welcome and Day 1 Recap Co-chairs
8:30 -9:30
‘White Space’ Discussion I
Discussion of topics which did not t into
the framework of day 1 but need to be
discussed.
9:30—10:30
‘White Space’ Discussion II
Discussion of parcularly far-out (or long-term),
high-risk, high-impact ideas.
10:30—10:45 BREAK
10:45—11:45 Discussion of Key Ideas/Components for Report
11:45—12:00 Closing Remarks Co-chairs
12:00 DEPARTURE

36
Future Directions Workshop: Advancing the Next Scientic Revolution in Toxicology
Basic Research Ofce, Ofce of the Under Secretary of Defense (RandE)
28-29 April 2022
Basic Research Innovative Collaboration Center
4100 N. Fairfax Road, Suite 450 Arlington, VA 22203
Co-Chairs: Thomas Hartung (Johns Hopkins), Ana Navas-Acien (Columbia), Weihsueh Chiu (Texas A&M)
In the nearly two decades since the human genome was sequenced, the eld of toxicology has undergone a transformation, taking
advantage of the explosion in biomedical knowledge and technologies to move from a largely empirical science aimed at ensuring
the absence of harmful effects to a mechanistic endeavor aimed at elucidating disease etiology. However, a substantial gap remains
between the promise of mechanistic toxicology and the actualization of the eld as a predictive science. For instance, high-throughput
in vitro and in silico toxicity testing remains largely focused on prioritization of individual chemicals for future investigation. Moreover,
efforts to translate such data into hazard or risk have been hampered by inadequate coverage of important biological targets,
inadequate consideration of population heterogeneity, and aiming still to provide assurances of safety rather than quantication of
effects across the population. Furthermore, there has been little progress on understanding the complex interactions among chemicals
and between chemicals and other intrinsic and extrinsic factors that affect population health, such as genetics and non-chemical
stressors, including marginalization and other social determinants of health.
This Future Directions Workshop on Advancing the Next Scientic Revolution in Toxicology aims to establish a new overarching vision
for toxicology as a predictive science. This vision entails a major paradigm shift in how toxicology is both conceived and practiced,
recognizing the multi-factorial, multi-causal nature of toxicity. Specically, this vision involves two critical steps:
• Moving away from reductionist interrogation of single chemicals, individual model systems, and discrete biological targets, which
ultimately cover only a minute sliver of relevant human experiences.
• Striving for a holistic understanding of the interactions among chemicals, non-chemical stressors, heterogeneous populations, and
life-stages, in order to prospectively identify and quantify their impacts on the incidence and severity of human disease.
A key outcome of this Workshop will be a roadmap of key basic science research needs that, if addressed in the next 10-20 years,
can substantially advance this transformational vision. The discussions and ensuing distributed report will provide valuable long-term
guidance to the DoD community, as well as the broader federal funding community, federal labs, and other stakeholders. Workshop
attendees will emerge with a better ability to identify and seize potential opportunities in the different elds addressed. This workshop
is sponsored by the Basic Research Ofce within the Ofce of Secretary of Defense, along with input and interest from the Services and
other DoD components.
Agenda
Rather than a standard conference format, the workshop design emphasizes interactive dialogue with primarily small-group breakout
sessions followed by whole-group synthesis of ideas.
Day One: The majority of the rst day will be spent in small-group breakout sessions on fundamental challenges to progress and
technical capabilities. The three breakout themes include:
1. Exposure-driven Toxicology
Populations are exposed to multiple environmental agents, including chemical agents through air, water, food and soil, and non-
chemical agents such as noise, light, and social stressors (e.g., racism, socioeconomic deprivation, climate). Toxicological research
that embraces an exposure-driven approach, characterizing real-life exposure scenarios including exposure mixtures and how these
agents work together affecting multiple mechanistic pathways and health outcomes is needed. A key opportunity is the expansion
of exposomic approaches to include this broader landscape of exposures. The interplay of environmental and social stressors
with genetic and molecular variants, and the contributions of this research towards the identication and evaluation of effective
interventions will be critical elements for discussion.
2. Technical Advances and Challenges
Predictive toxicology requires expanding the “toolbox” in several directions. First, because adverse outcomes involve interactions
of environment (see above), genes, and lifestage, we need our “model systems” to cover “gene” and “lifestage” more broadly.
37
Example technologies include genetically diverse population-based in vitro and in vivo resources, and expansion of experimental
designs to cover different stages of development, as well as developmental origins of health and disease. Additionally, our
approaches currently cluster at the beginning (e.g., high throughput assays) and the end (e.g., in vivo apical endpoints) of the
pathophysiological process, neglecting the modulating and stochastic factors that inuence outcomes that lie between. Thus,
approaches that provide access to intermediate states, perturbations, and outcomes are needed to better understand the
progression to disease. Example technologies include novel biomarkers, microphysiological systems (e.g., organ on a chip), and
in silico models (e.g., systems toxicology/virtual experiments, AI/Machine Learning). Finally, a key challenge is characterizing the
predictive accuracy, precision, and relevance of new approaches, as well as understanding their domains of applicability.
3. Evidence Integration
Toxicology is currently transitioning from a data-poor to a data-rich science with the curation of legacy databases, “grey”
information in the internet, mining of scientic literature, sensor technologies, ~omics, robotized testing, high-content imaging
and others. Key questions include how to handle these new types of information sources, which may be incomplete, how to weigh
(evidence strength, risk of bias, quality scoring etc.), and how to integrate this evidence. For instance, probabilistic risk assessment
integrates across sources of evidence resulting in a more holistic probability of risk/hazard, though challenges include how to
validate (real-life, t for purpose, ground truthing, qualication, triangulation) and communicate these probabilities. Additional
challenges, such as data curation and storage, mining, analysis and visualization will be discussed.
Day Two: The second day of the workshop is a half-day consisting of white-space, whole group discussions on topics that did not
fall into the Day 1 framework or were especially ambitious and/or high-risk. Participants will also discuss cross cutting themes and
the trajectory of the eld over the next 10-20 years. At the end of the day, the whole group will discuss the overarching themes of the
workshop that should be included in the nal workshop report.
Cross Cutting themes to discuss
• What disease endpoints are the most promising in terms of developing the knowledge (e.g., availability of human biomarkers,
understanding of genetic/non-genetic risk factors) and technologies (e.g., microphysiological systems, computational models)
needed to enable predictive toxicology?
• Where are the greatest opportunities for synergies between toxicology and other disciplines including the social sciences?
• How to ensure standards (e.g., reporting standards, systematic review, meta-analysis, risk of bias analysis, etc.) that retain quality
assurance (best practices, validation and other aspects of QA and QC) and public health protection in a mechanistic toxicology?
